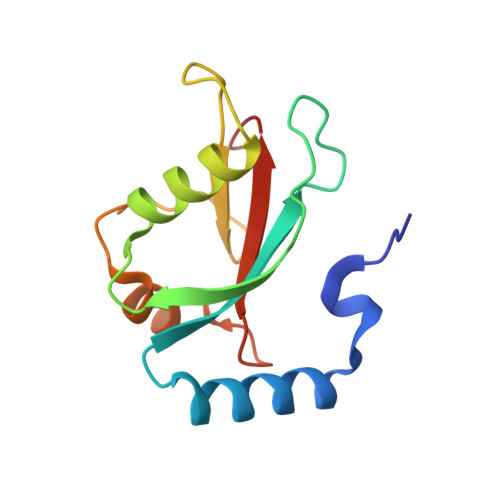
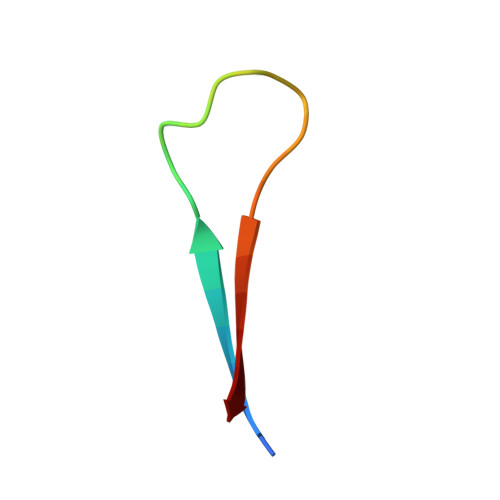
A novel conformation of the LC3-interacting region motif revealed by the structure of a complex between LC3B and RavZ
Kwon, D.H., Kim, L., Kim, B.W., Kim, J.H., Roh, K.H., Choi, E.J., Song, H.K.(2017) Biochem Biophys Res Commun 490: 1093-1099
- PubMed: 28668392
- DOI: https://doi.org/10.1016/j.bbrc.2017.06.173
- Primary Citation Related Structures:
5XAC, 5XAD, 5XAE - PubMed Abstract:
LC3-family member proteins play a critical role in autophagy, a cellular process responsible for the degradation of massive cellular components including intracellular pathogens. A variety of molecules involved in the autophagic pathway engage in specific interactions with a unique sequence motif referred to as the LIR (LC3-interacting region) motif. Although identification of conserved structural features of LIR motifs in complex with LC3-family members has established a canonical LIR motif, atypical conformations of LIR motifs have recently been revealed. Here, we determined the three-dimensional crystal structures of LC3B in complex with three different LIR motifs of RavZ from Legionella pneumophila, an intracellular pathogen that can manipulate the host autophagy system. The tandem LIR motifs located in the N-terminal region of RavZ adopt a novel β-sheet conformation and thus provide specific ionic interactions with LC3B in addition to canonical hydrophobic plugged-in interactions. Consequently, these motifs possess higher binding affinity to LC3-family members than canonical LIR motifs, although the tandem repeats can only bind to one LC3 molecule. These findings broaden our understanding of the functional repertoire of LIR motifs in autophagy.
- Department of Life Sciences, Korea University, 145 Anam-ro, Seongbuk-gu, Seoul 02841, Republic of Korea.
Organizational Affiliation: