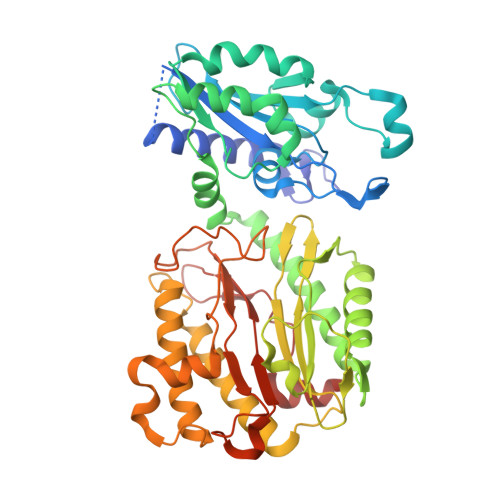
Structure of the human aminopeptidase XPNPEP3 and comparison of its in vitro activity with Icp55 orthologs: Insights into diverse cellular processes.
Singh, R., Jamdar, S.N., Goyal, V.D., Kumar, A., Ghosh, B., Makde, R.D.(2017) J Biological Chem 292: 10035-10047
- PubMed: 28476889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.783357
- Primary Citation Related Structures:
5X49 - PubMed Abstract:
The human aminopeptidase XPNPEP3 is associated with cystic kidney disease and TNF-TNFR2 cellular signaling. Its yeast and plant homolog Icp55 processes several imported mitochondrial matrix proteins leading to their stabilization. However, the molecular basis for the diverse roles of these enzymes in the cell is unknown. Here, we report the crystal structure of human XPNPEP3 with bound apstatin product at 1.65 Å resolution, and we compare its in vitro substrate specificity with those of fungal Icp55 enzymes. In contrast to the suggestions by earlier in vivo studies of mitochondrial processing, we found that these enzymes are genuine Xaa-Pro aminopeptidases, which hydrolyze peptides with proline at the second position (P1'). The mitochondrial processing activity involving cleavage of peptides lacking P1' proline was also detected in the purified enzymes. A wide proline pocket as well as molecular complementarity and capping at the S1 substrate site of XPNPEP3 provide the necessary structural features for processing the mitochondrial substrates. However, this activity was found to be significantly lower as compared with Xaa-Pro aminopeptidase activity. Because of similar activity profiles of Icp55 and XPNPEP3, we propose that XPNPEP3 plays the same mitochondrial role in humans as Icp55 does in yeast. Both Xaa-Pro aminopeptidase and mitochondrial processing activities of XPNPEP3 have implications toward mitochondrial fitness and cystic kidney disease. Furthermore, the presence of both these activities in Icp55 elucidates the unexplained processing of the mitochondrial cysteine desulfurase Nfs1 in yeast. The enzymatic and structural analyses reported here provide a valuable molecular framework for understanding the diverse cellular roles of XPNPEP3.
- From the High Pressure and Synchrotron Radiation Physics Division and.
Organizational Affiliation: