Structure of p-Ub-S65-NH2 at 1.82 Angstroms resolution
Man, P., Shuai, G., Qian, Q., Lei, L.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Polyubiquitin-B | 76 | Homo sapiens | Mutation(s): 0 | 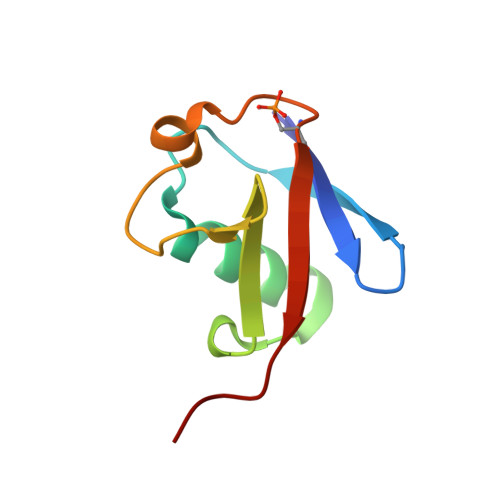 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P0CG47 (Homo sapiens) Explore P0CG47 Go to UniProtKB: P0CG47 | |||||
PHAROS: P0CG47 GTEx: ENSG00000170315 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0CG47 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| D-ubiquitin | B [auth D] | 76 | Homo sapiens | Mutation(s): 0 | 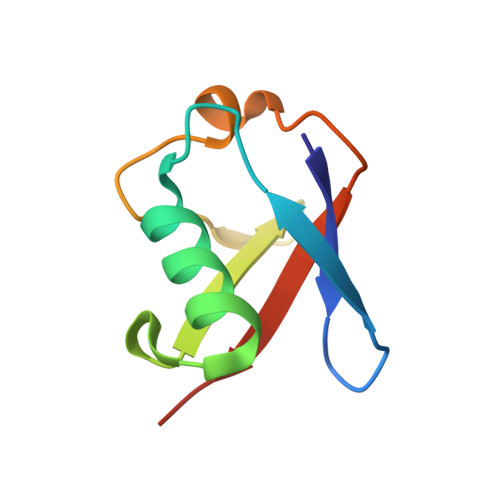 |
Sequence AnnotationsExpand | |||||
| |||||
| Modified Residues 18 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| SEP Query on SEP | A | L-PEPTIDE LINKING | C3 H8 N O6 P |  | SER |
| DAL Query on DAL | B [auth D] | D-PEPTIDE LINKING | C3 H7 N O2 |  | -- |
| DAR Query on DAR | B [auth D] | D-PEPTIDE LINKING | C6 H15 N4 O2 |  | -- |
| DAS Query on DAS | B [auth D] | D-PEPTIDE LINKING | C4 H7 N O4 |  | -- |
| DGL Query on DGL | B [auth D] | D-PEPTIDE LINKING | C5 H9 N O4 |  | -- |
| DGN Query on DGN | B [auth D] | D-PEPTIDE LINKING | C5 H10 N2 O3 |  | -- |
| DHI Query on DHI | B [auth D] | D-PEPTIDE LINKING | C6 H10 N3 O2 |  | -- |
| DIL Query on DIL | B [auth D] | D-PEPTIDE LINKING | C6 H13 N O2 |  | -- |
| DLE Query on DLE | B [auth D] | D-PEPTIDE LINKING | C6 H13 N O2 |  | -- |
| DLY Query on DLY | B [auth D] | D-PEPTIDE LINKING | C6 H14 N2 O2 |  | -- |
| DNE Query on DNE | B [auth D] | D-PEPTIDE LINKING | C6 H13 N O2 |  | -- |
| DPN Query on DPN | B [auth D] | D-PEPTIDE LINKING | C9 H11 N O2 |  | -- |
| DPR Query on DPR | B [auth D] | D-PEPTIDE LINKING | C5 H9 N O2 |  | -- |
| DSG Query on DSG | B [auth D] | D-PEPTIDE LINKING | C4 H8 N2 O3 |  | -- |
| DSN Query on DSN | B [auth D] | D-PEPTIDE LINKING | C3 H7 N O3 |  | -- |
| DTH Query on DTH | B [auth D] | D-PEPTIDE LINKING | C4 H9 N O3 |  | -- |
| DTY Query on DTY | B [auth D] | D-PEPTIDE LINKING | C9 H11 N O3 |  | -- |
| DVA Query on DVA | B [auth D] | D-PEPTIDE LINKING | C5 H11 N O2 |  | -- |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 28.914 | α = 81.91 |
| b = 29.969 | β = 72.38 |
| c = 39.726 | γ = 62.43 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data reduction |