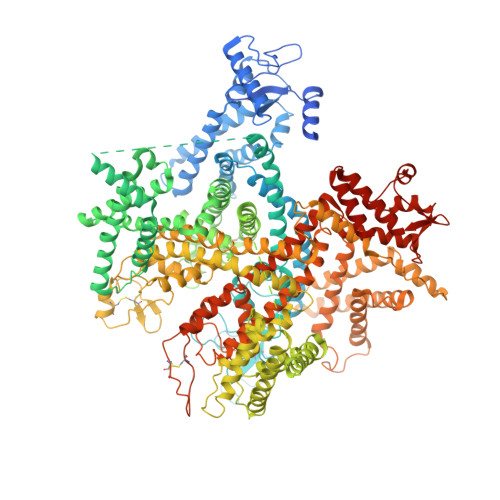
Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution.
Shen, H., Zhou, Q., Pan, X., Li, Z., Wu, J., Yan, N.(2017) Science 355
- PubMed: 28183995 Search on PubMed
- DOI: https://doi.org/10.1126/science.aal4326
- Primary Citation Related Structures:
5X0M - PubMed Abstract:
Voltage-gated sodium (Na v ) channels are responsible for the initiation and propagation of action potentials. They are associated with a variety of channelopathies and are targeted by multiple pharmaceutical drugs and natural toxins. Here, we report the cryogenic electron microscopy structure of a putative Na v channel from American cockroach (designated Na v PaS) at 3.8 angstrom resolution. The voltage-sensing domains (VSDs) of the four repeats exhibit distinct conformations. The entrance to the asymmetric selectivity filter vestibule is guarded by heavily glycosylated and disulfide bond-stabilized extracellular loops. On the cytoplasmic side, a conserved amino-terminal domain is placed below VSD I , and a carboxy-terminal domain binds to the III-IV linker. The structure of Na v PaS establishes an important foundation for understanding function and disease mechanism of Na v and related voltage-gated calcium channels.
- State Key Laboratory of Membrane Biology, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: