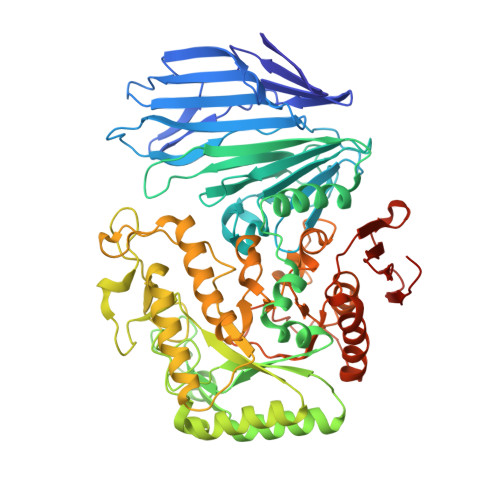
The first crystal structure of a family 129 glycoside hydrolase from a probiotic bacterium reveals critical residues and metal cofactors
Sato, M., Liebschner, D., Yamada, Y., Matsugaki, N., Arakawa, T., Wills, S.S., Hattie, M., Stubbs, K.A., Ito, T., Senda, T., Ashida, H., Fushinobu, S.(2017) J Biological Chem 292: 12126-12138
- PubMed: 28546425
- DOI: https://doi.org/10.1074/jbc.M117.777391
- Primary Citation of Related Structures:
5WZN, 5WZP, 5WZQ, 5WZR - PubMed Abstract:
The α- N -acetylgalactosaminidase from the probiotic bacterium Bifidobacterium bifidum (NagBb) belongs to the glycoside hydrolase family 129 and hydrolyzes the glycosidic bond of Tn-antigen (GalNAcα1-Ser/Thr). NagBb is involved in assimilation of O -glycans on mucin glycoproteins by B. bifidum in the human gastrointestinal tract, but its catalytic mechanism has remained elusive because of a lack of sequence homology around putative catalytic residues and of other structural information. Here we report the X-ray crystal structure of NagBb, representing the first GH129 family structure, solved by the single-wavelength anomalous dispersion method based on sulfur atoms of the native protein. We determined ligand-free, GalNAc, and inhibitor complex forms of NagBb and found that Asp-435 and Glu-478 are located in the catalytic domain at appropriate positions for direct nucleophilic attack at the anomeric carbon and proton donation for the glycosidic bond oxygen, respectively. A highly conserved Asp-330 forms a hydrogen bond with the O4 hydroxyl of GalNAc in the -1 subsite, and Trp-398 provides a stacking platform for the GalNAc pyranose ring. Interestingly, a metal ion, presumably Ca 2+ , is involved in the recognition of the GalNAc N -acetyl group. Mutations at Asp-435, Glu-478, Asp-330, and Trp-398 and residues involved in metal coordination (including an all-Ala quadruple mutant) significantly reduced the activity, indicating that these residues and the metal ion play important roles in substrate recognition and catalysis. Interestingly, NagBb exhibited some structural similarities to the GH101 endo-α- N -acetylgalactosaminidases, but several critical differences in substrate recognition and reaction mechanism account for the different activities of these two enzymes.
- Department of Biotechnology, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: