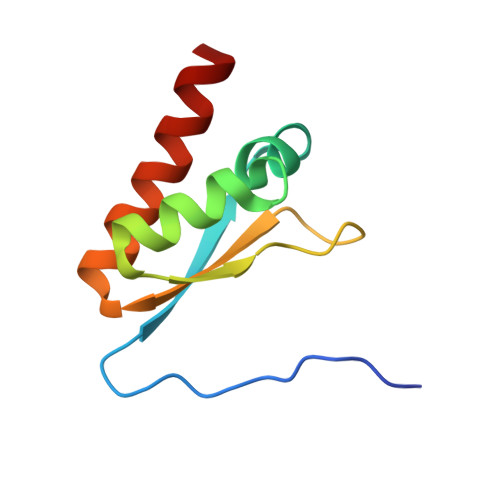
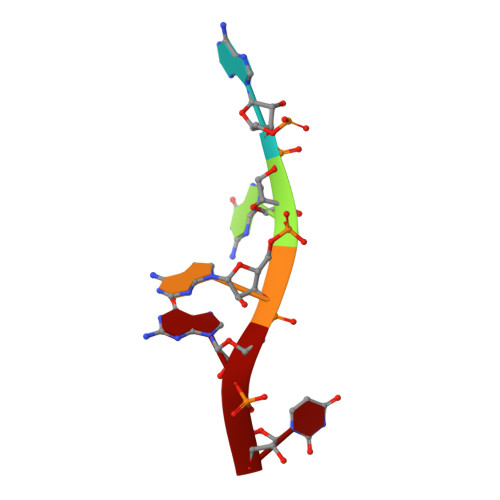
The human RNA-binding protein and E3 ligase MEX-3C binds the MEX-3-recognition element (MRE) motif with high affinity
Yang, L., Wang, C., Li, F., Zhang, J., Nayab, A., Wu, J., Shi, Y., Gong, Q.(2017) J Biological Chem 292: 16221-16234
- PubMed: 28808060 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.797746
- Primary Citation Related Structures:
5WWW, 5WWX, 5WWZ - PubMed Abstract:
MEX-3 is a K-homology (KH) domain-containing RNA-binding protein first identified as a translational repressor in Caenorhabditis elegans , and its four orthologs (MEX-3A-D) in human and mouse were subsequently found to have E3 ubiquitin ligase activity mediated by a RING domain and critical for RNA degradation. Current evidence implicates human MEX-3C in many essential biological processes and suggests a strong connection with immune diseases and carcinogenesis. The highly conserved dual KH domains in MEX-3 proteins enable RNA binding and are essential for the recognition of the 3'-UTR and post-transcriptional regulation of MEX-3 target transcripts. However, the molecular mechanisms of translational repression and the consensus RNA sequence recognized by the MEX-3C KH domain are unknown. Here, using X-ray crystallography and isothermal titration calorimetry, we investigated the RNA-binding activity and selectivity of human MEX-3C dual KH domains. Our high-resolution crystal structures of individual KH domains complexed with a noncanonical U-rich and a GA-rich RNA sequence revealed that the KH1/2 domains of human MEX-3C bound MRE10, a 10-mer RNA (5'-CAGAGUUUAG-3') consisting of an eight-nucleotide MEX-3-recognition element (MRE) motif, with high affinity. Of note, we also identified a consensus RNA motif recognized by human MEX-3C. The potential RNA-binding sites in the 3'-UTR of the human leukocyte antigen serotype ( HLA-A2 ) mRNA were mapped with this RNA-binding motif and further confirmed by fluorescence polarization. The binding motif identified here will provide valuable information for future investigations of the functional pathways controlled by human MEX-3C and for predicting potential mRNAs regulated by this enzyme.
- From the Hefei National Laboratory for Physical Science at Microscale, Collaborative Innovation Center of Chemistry for Life Sciences and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230027, China and.
Organizational Affiliation: