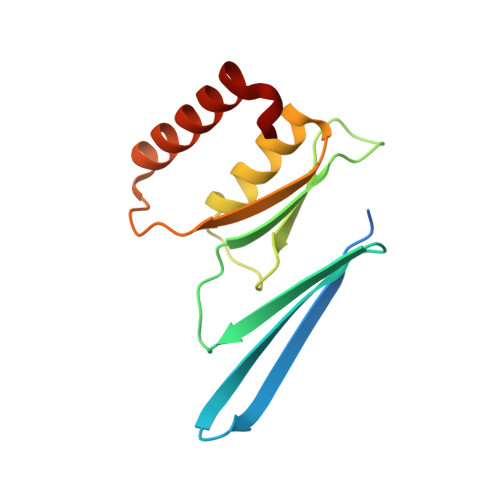
Crystal structure of human proteasome assembly chaperone PAC4 involved in proteasome formation
Kurimoto, E., Satoh, T., Ito, Y., Ishihara, E., Okamoto, K., Yagi-Utsumi, M., Tanaka, K., Kato, K.(2017) Protein Sci 26: 1080-1085
- PubMed: 28263418 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3153
- Primary Citation Related Structures:
5WTQ - PubMed Abstract:
The 26S proteasome is a large protein complex, responsible for degradation of ubiquinated proteins in eukaryotic cells. Eukaryotic proteasome formation is a highly ordered process that is assisted by several assembly chaperones. The assembly of its catalytic 20S core particle depends on at least five proteasome-specific chaperones, i.e., proteasome-assembling chaperons 1-4 (PAC1-4) and proteasome maturation protein (POMP). The orthologues of yeast assembly chaperones have been structurally characterized, whereas most mammalian assembly chaperones are not. In the present study, we determined a crystal structure of human PAC4 at 1.90-Å resolution. Our crystallographic data identify a hydrophobic surface that is surrounded by charged residues. The hydrophobic surface is complementary to that of its binding partner, PAC3. The surface also exhibits charge complementarity with the proteasomal α4-5 subunits. This will provide insights into human proteasome-assembling chaperones as potential anticancer drug targets.
- Faculty of Pharmacy, Meijo University, Tempaku-ku, Nagoya, 468-8503, Japan.
Organizational Affiliation: