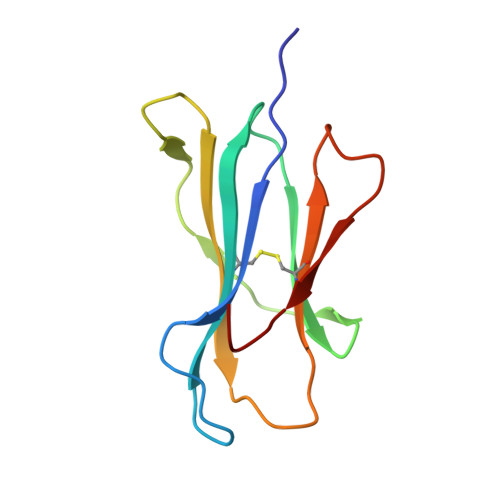
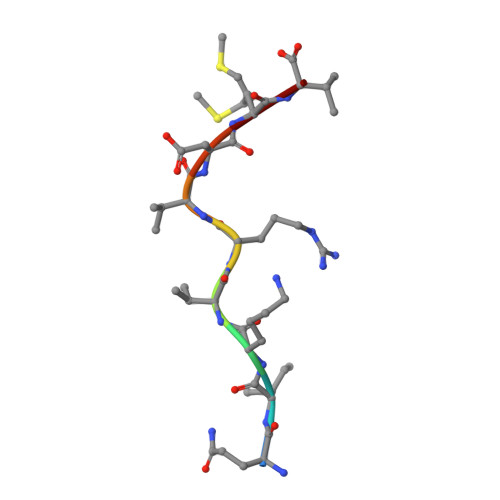
Inability To Detect Cross-Reactive Memory T Cells Challenges the Frequency of Heterologous Immunity among Common Viruses.
Rowntree, L.C., Nguyen, T.H.O., Halim, H., Purcell, A.W., Rossjohn, J., Gras, S., Kotsimbos, T.C., Mifsud, N.A.(2018) J Immunol 200: 3993-4003
- PubMed: 29735483 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.1800010
- Primary Citation Related Structures:
5WMN, 5WMO, 5WMP, 5WMQ, 5WMR - PubMed Abstract:
Human memory T cells that cross-react with epitopes from unrelated viruses can potentially modulate immune responses to subsequent infections by a phenomenon termed heterologous immunity. However, it is unclear whether similarities in structure rather than sequence underpin heterologous T cell cross-reactivity. In this study, we aimed to explore the mechanism of heterologous immunity involving immunodominant epitopes derived from common viruses restricted to high-frequency HLA allotypes (HLA-A*02:01, -B*07:02, and -B*08:01). We examined EBV-specific memory T cells for their ability to cross-react with CMV or influenza A virus-derived epitopes. Following T cell immunoassays to determine phenotype and function, complemented with biophysical and structural investigations of peptide/HLA complexes, we did not detect cross-reactivity of EBV-specific memory T cells toward either CMV or influenza A virus epitopes presented by any of the selected HLA allomorphs. Thus, despite the ubiquitous nature of these human viruses and the dominant immune response directed toward the selected epitopes, heterologous virus-specific T cell cross-reactivity was not detected. This suggests that either heterologous immunity is not as common as previously reported, or that it requires a very specific biological context to develop and be clinically relevant.
- Department of Medicine, Monash University, Central Clinical School, The Alfred Hospital, Melbourne, Victoria 3004, Australia.
Organizational Affiliation: