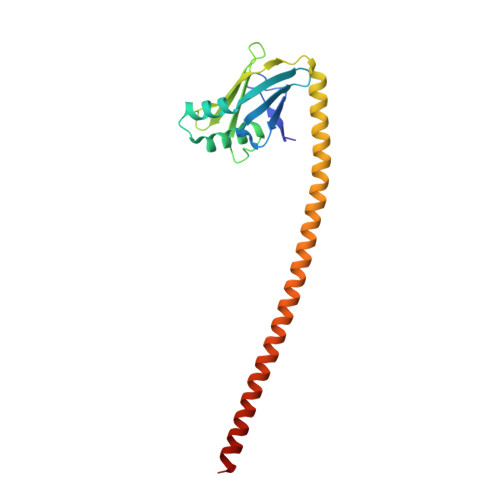
Design considerations in coiled-coil fusion constructs for the structural determination of a problematic region of the human cardiac myosin rod.
Andreas, M.P., Ajay, G., Gellings, J.A., Rayment, I.(2017) J Struct Biol 200: 219-228
- PubMed: 28743637 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2017.07.006
- Primary Citation Related Structures:
5WJ7, 5WJB, 5WLQ, 5WLZ, 5WME - PubMed Abstract:
X-ray structural determination of segments of the myosin rod has proved difficult because of the strong salt-dependent aggregation properties and repeating pattern of charges on the surface of the coiled-coil that lead to the formation of paracrystals. This problem has been resolved in part through the use of globular assembly domains that improve protein folding and prevent aggregation. The primary consideration now in designing coiled-coil fusion constructs for myosin is deciding where to truncate the coiled-coil and which amino acid residues to include from the folding domain. This is especially important for myosin that contains numerous regions of low predicted coiled-coil propensity. Here we describe the strategy adopted to determine the structure of the region that extends from Arg1677 - Leu1797 that included two areas that do not show a strong sequence signature of a conventional left-handed coiled coil or canonical heptad repeat. This demonstrates again that, with careful choice of fusion constructs, overlapping structures exhibit very similar conformations for the myosin rod fragments in the canonical regions. However, conformational variability is seen around Leu1706 which is a hot spot for cardiomyopathy mutations suggesting that this might be important for function.
- Department of Biochemistry, University of Wisconsin-Madison, WI 53706, USA.
Organizational Affiliation: