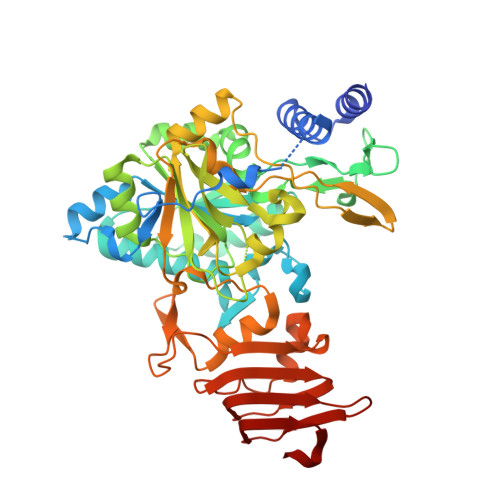
Crystal structure and insights into the oligomeric state of UDP-glucose pyrophosphorylase from sugarcane.
Cotrim, C.A., Soares, J.S.M., Kobe, B., Menossi, M.(2018) PLoS One 13: e0193667-e0193667
- PubMed: 29494650 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0193667
- Primary Citation Related Structures:
5WEG - PubMed Abstract:
UDP-glucose pyrophosphorylase (UGPase) is found in all organisms and catalyses the formation of UDP-glucose. In sugarcane, UDP-glucose is a branch-point in the carbon channelling into other carbohydrates, such as sucrose and cellulose, which are the major factors for sugarcane productivity. In most plants, UGPase has been described to be enzymatically active in the monomeric form, while in human and yeast, homo-octamers represent the active form of the protein. Here, we present the crystal structure of UGPase from sugarcane (ScUGPase-1) at resolution of 2.0 Å. The crystals of ScUGPase-1 reveal the presence of two molecules in the asymmetric unit and the multi-angle light scattering analysis shows that ScUGPase-1 forms a mixture of species ranging from monomers to larger oligomers in solution, suggesting similarities with the orthologs from yeast and human.
- School of Chemistry and Molecular Bioscience and Australia Infectious Diseases Research Centre, University of Queensland, Brisbane, Queensland, Australia.
Organizational Affiliation: