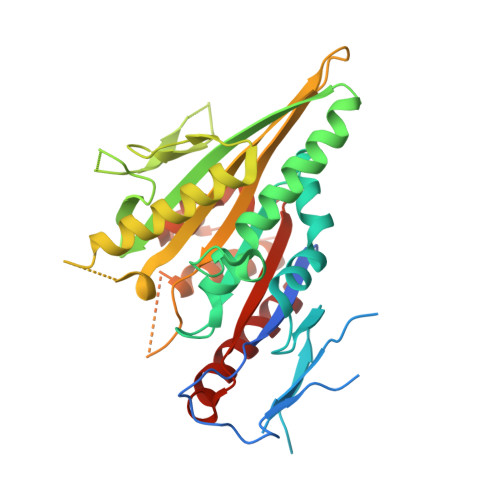
Structural basis of small molecule ATPase inhibition of a human mitotic kinesin motor protein.
Park, H.W., Ma, Z., Zhu, H., Jiang, S., Robinson, R.C., Endow, S.A.(2017) Sci Rep 7: 15121-15121
- PubMed: 29123223 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-14754-6
- Primary Citation Related Structures:
5W3D, 5WDE, 5WDH - PubMed Abstract:
Kinesin microtubule motor proteins play essential roles in division, including attaching chromosomes to spindles and crosslinking microtubules for spindle assembly. Human kinesin-14 KIFC1 is unique in that cancer cells with amplified centrosomes are dependent on the motor for viable division because of its ability to cluster centrosomes and form bipolar spindles, but it is not required for division in almost all normal cells. Screens for small molecule inhibitors of KIFC1 have yielded several candidates for further development, but obtaining structural data to determine their sites of binding has been difficult. Here we compare a previously unreported KIFC1 crystal structure with new structures of two closely related kinesin-14 proteins, Ncd and KIFC3, to determine the potential binding site of a known KIFC1 ATPase inhibitor, AZ82. We analyze the previously identified kinesin inhibitor binding sites and identify features of AZ82 that favor binding to one of the sites, the α4/α6 site. This selectivity can be explained by unique structural features of the KIFC1 α4/α6 binding site. These features may help improve the drug-like properties of AZ82 and other specific KIFC1 inhibitors.
- Structural Genomics Consortium, University of Toronto, Toronto, ON M5G 1L7, Canada.
Organizational Affiliation: