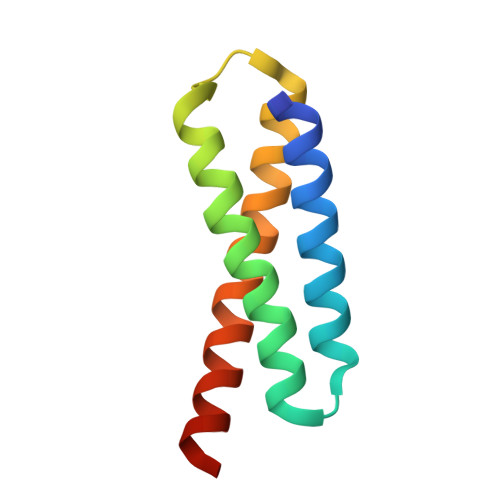
Crystal structure of the Legionella effector Lem22.
Kozlov, G., Wong, K., Gehring, K.(2018) Proteins 86: 263-267
- PubMed: 29159828 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.25427
- Primary Citation Related Structures:
5WD8, 5WD9 - PubMed Abstract:
Legionella pneumophila is a pathogen causing severe pneumonia in humans called Legionnaires' disease. Lem22 is a previously uncharacterized effector protein conserved in multiple Legionella strains. Here, we report the crystal structure of Lem22 from the Philadelphia strain, also known as lpg2328, at 1.40 Å resolution. The structure shows an up-and-down three-helical bundle with a significant structural similarity to a number of protein-binding domains involved in apoptosis and membrane trafficking. Sequence conservation identifies a putative functional site on the interface of helices 2 and 3. The structure is an important step toward a functional characterization of Lem22.
- Department of Biochemistry, Groupe de recherche axé sur la structure des protéines, McGill University, Montreal, QC, H3G 0B1, Canada.
Organizational Affiliation: