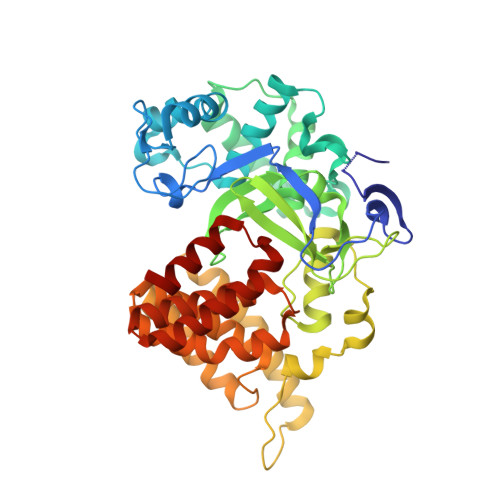
The crystal structure of SMYD2 in complex with compound MTF003
Zeng, H., Dong, A., Walker, J.R., Hutch, A., Seitova, A., Tatlock, J., Kumpf, R., Owen, A., Taylor, A., Casimiro-Garcia, A., Bountra, C., Arrowsmith, C.H., Edwards, A.M., Brown, P.J., Wu, H.To be published.