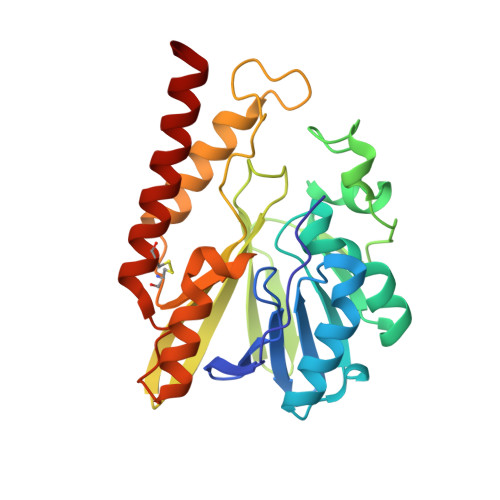
Three-dimensional structure of FEZ-1, a monomeric subclass B3 metallo-beta-lactamase from Fluoribacter gormanii, in native form and in complex with D-captopril.
Garcia-Saez, I., Mercuri, P.S., Papamicael, C., Kahn, R., Frere, J.M., Galleni, M., Rossolini, G.M., Dideberg, O.(2003) J Mol Biology 325: 651-660
- PubMed: 12507470 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)01271-8
- Primary Citation Related Structures:
1L9Y, 5W90, 5WCK - PubMed Abstract:
The beta-lactamases are involved in bacterial resistance to penicillin and related compounds. Members of the metallo-enzyme class are now found in many pathogenic bacteria and are thus becoming of major clinical importance. The structures of the Zn-beta-lactamase from Fluoribacter gormanii (FEZ-1) in the native and in the complex form are reported here. FEZ-1 is a monomeric enzyme, which possesses two zinc-binding sites. These structures are discussed in comparison with those of the tetrameric L1 enzyme produced by Stenotrophomonas maltophilia. From this analysis, amino acids involved in the oligomerization of L1 are clearly identified. Despite the similarity in fold, the active site of FEZ-1 was found to be significantly different. Two residues, which were previously implicated in function, are not present in L1 or in FEZ-1. The broad-spectrum substrate profile of Zn-beta-lactamases arises from the rather wide active-site cleft, where various beta-lactam compounds can be accommodated.
- Institut de Biologie Structurale (CIP), Jean-Pierre Ebel (CNRS-CEA), 41 rue Jules Horowitz, 38027 Grenoble Cedex 1, France.
Organizational Affiliation: