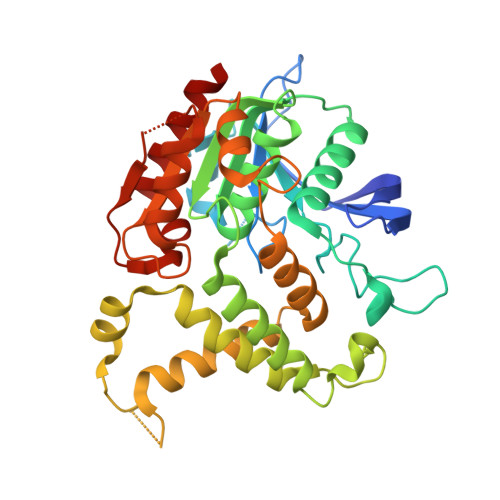
Structural analysis of mycobacterial homoserine transacetylases central to methionine biosynthesis reveals druggable active site.
Chaton, C.T., Rodriguez, E.S., Reed, R.W., Li, J., Kenner, C.W., Korotkov, K.V.(2019) Sci Rep 9: 20267-20267
- PubMed: 31889085 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-56722-2
- Primary Citation Related Structures:
5W8O, 5W8P, 6PUX - PubMed Abstract:
Mycobacterium tuberculosis is the cause of the world's most deadly infectious disease. Efforts are underway to target the methionine biosynthesis pathway, as it is not part of the host metabolism. The homoserine transacetylase MetX converts L-homoserine to O-acetyl-L-homoserine at the committed step of this pathway. In order to facilitate structure-based drug design, we determined the high-resolution crystal structures of three MetX proteins, including M. tuberculosis (MtMetX), Mycolicibacterium abscessus (MaMetX), and Mycolicibacterium hassiacum (MhMetX). A comparison of homoserine transacetylases from other bacterial and fungal species reveals a high degree of structural conservation amongst the enzymes. Utilizing homologous structures with bound cofactors, we analyzed the potential ligandability of MetX. The deep active-site tunnel surrounding the catalytic serine yielded many consensus clusters during mapping, suggesting that MtMetX is highly druggable.
- Department of Molecular & Cellular Biochemistry and the Center for Structural Biology, University of Kentucky, Lexington, KY, 40536, USA.
Organizational Affiliation: