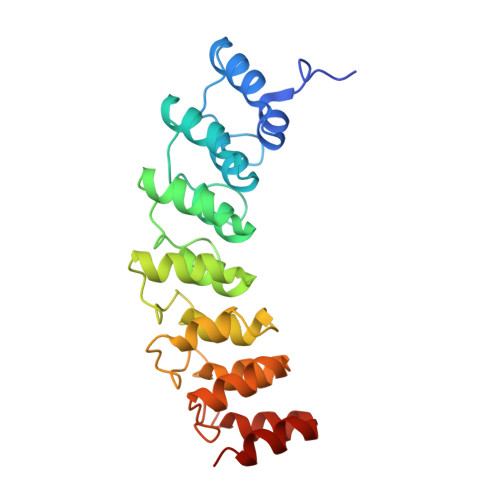
Structural Basis for Substrate Recognition by the Ankyrin Repeat Domain of Human DHHC17 Palmitoyltransferase.
Verardi, R., Kim, J.S., Ghirlando, R., Banerjee, A.(2017) Structure 25: 1337-1347.e6
- PubMed: 28757145 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2017.06.018
- Primary Citation Related Structures:
5W7I, 5W7J - PubMed Abstract:
DHHC enzymes catalyze palmitoylation, a major post-translational modification that regulates a number of key cellular processes. There are up to 24 DHHCs in mammals and hundreds of substrate proteins that get palmitoylated. However, how DHHC enzymes engage with their substrates is still poorly understood. There is currently no structural information about the interaction between any DHHC enzyme and protein substrates. In this study we have investigated the structural and thermodynamic bases of interaction between the ankyrin repeat domain of human DHHC17 (ANK17) and Snap25b. We solved a high-resolution crystal structure of the complex between ANK17 and a peptide fragment of Snap25b. Through structure-guided mutagenesis, we discovered key residues in DHHC17 that are critically important for interaction with Snap25b. We further extended our finding by showing that the same residues are also crucial for the interaction of DHHC17 with Huntingtin, one of its most physiologically relevant substrates.
- Unit on Structural and Chemical Biology of Membrane Proteins, Cell Biology and Neurobiology Branch, National Institute of Child Health and Human Development, National Institutes of Health, 35A Convent Drive, Bethesda, MD 20892, USA.
Organizational Affiliation: