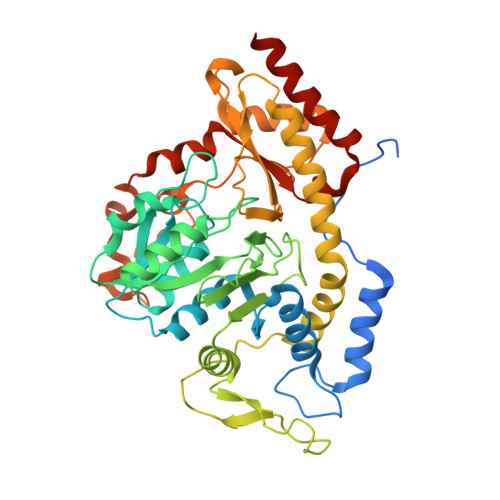
The structure of RbmB from Streptomyces ribosidificus, an aminotransferase involved in the biosynthesis of ribostamycin.
Zachman-Brockmeyer, T.R., Thoden, J.B., Holden, H.M.(2017) Protein Sci 26: 1886-1892
- PubMed: 28685903 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3221
- Primary Citation Related Structures:
5W70, 5W71 - PubMed Abstract:
Aminoglycoside antibiotics represent a classical group of antimicrobials first discovered in the 1940s. Due to their ototoxic and nephrotoxic side effects, they are typically only used against Gram negative bacteria which have become resistant to other therapeutics. One family of aminoglycosides includes such compounds as butirosin, ribostamycin, neomycin, and kanamycin, amongst others. The common theme in these antibiotics is that they are constructed around a chemically stable aminocyclitol unit referred to as 2-deoxystreptamine (2-DOS). Four enzymes are required for the in vivo production of 2-DOS. Here, we report the structure of RbmB from Streptomyces ribosidificus, which is a pyridoxal 5'-phosphate dependent enzyme that catalyzes two of the required steps in 2-DOS formation by functioning on distinct substrates. For this analysis, the structure of the external aldimine form of RbmB with 2-DOS was determined to 2.1 Å resolution. In addition, the structure of a similar enzyme, BtrR from Bacillus circulans, was also determined to 2.1 Å resolution in the same external aldimine form. These two structures represent the first detailed molecular descriptions of the active sites for those aminotransferases involved in 2-DOS production. Given the fact that the 2-DOS unit is widespread amongst aminoglycoside antibiotics, the data presented herein provide new molecular insight into the biosynthesis of these sugar-based drugs.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin.
Organizational Affiliation: