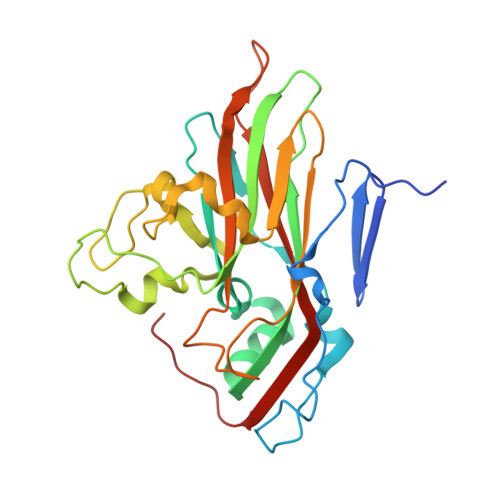
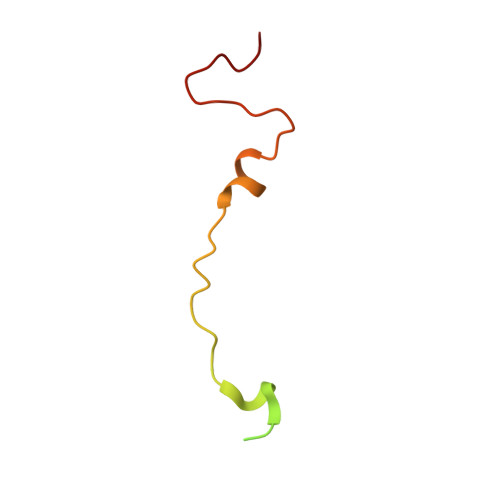
Antibody-induced uncoating of human rhinovirus B14.
Dong, Y., Liu, Y., Jiang, W., Smith, T.J., Xu, Z., Rossmann, M.G.(2017) Proc Natl Acad Sci U S A 114: 8017-8022
- PubMed: 28696310 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1707369114
- Primary Citation Related Structures:
5W3E, 5W3L, 5W3M, 5W3O - PubMed Abstract:
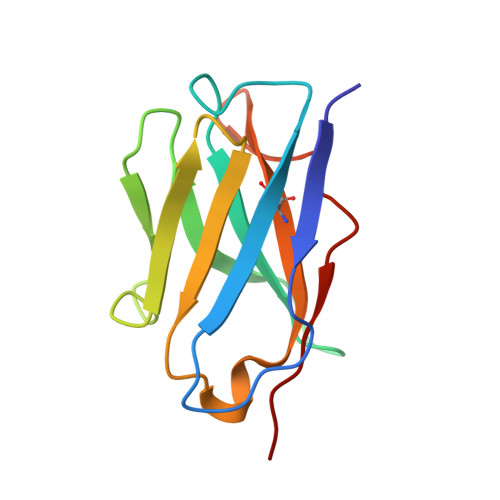
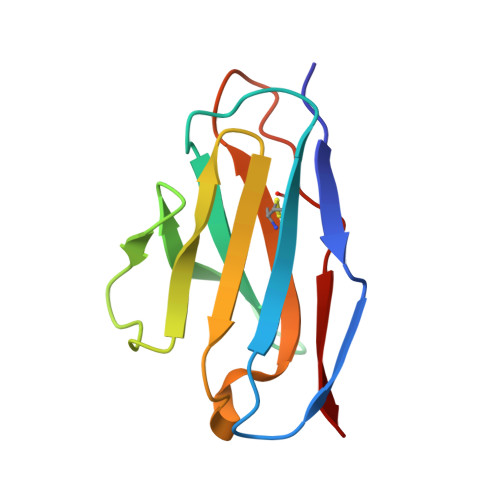
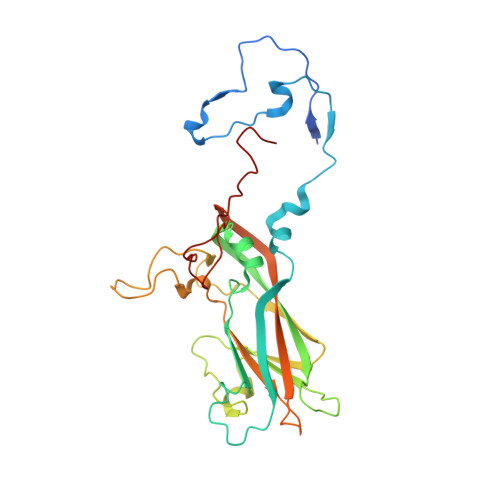
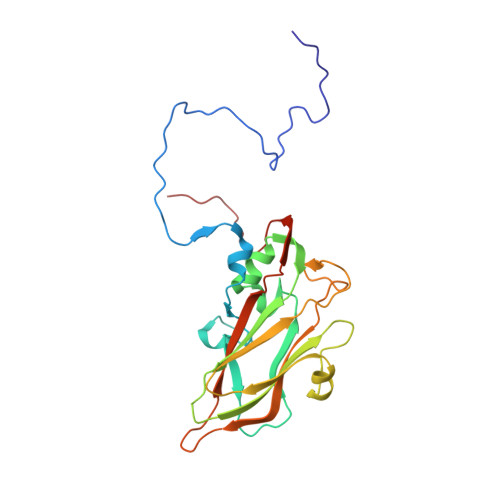
Rhinoviruses (RVs) are the major causes of common colds in humans. They have a nonenveloped, icosahedral capsid surrounding a positive-strand RNA genome. Here we report that the antigen-binding (Fab) fragment of a neutralizing antibody (C5) can trigger genome release from RV-B14 to form emptied particles and neutralize virus infection. Using cryo-electron microscopy, structures of the C5 Fab in complex with the full and emptied particles have been determined at 2.3 Å and 3.0 Å resolution, respectively. Each of the 60 Fab molecules binds primarily to a region on viral protein 3 (VP3). Binding of the C5 Fabs to RV-B14 results in significant conformational changes around holes in the capsid through which the viral RNA might exit. These results are so far the highest resolution view of an antibody-virus complex and elucidate a mechanism whereby antibodies neutralize RVs and related viruses by inducing virus uncoating.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907.
Organizational Affiliation: