A conserved energetic footprint underpins recognition of human leukocyte antigen-E by two distinct alpha beta T cell receptors.
Sullivan, L.C., Walpole, N.G., Farenc, C., Pietra, G., Sum, M.J.W., Clements, C.S., Lee, E.J., Beddoe, T., Falco, M., Mingari, M.C., Moretta, L., Gras, S., Rossjohn, J., Brooks, A.G.(2017) J Biological Chem 292: 21149-21158
- PubMed: 28972140
- DOI: https://doi.org/10.1074/jbc.M117.807719
- Primary Citation Related Structures:
5W1V, 5W1W - PubMed Abstract:
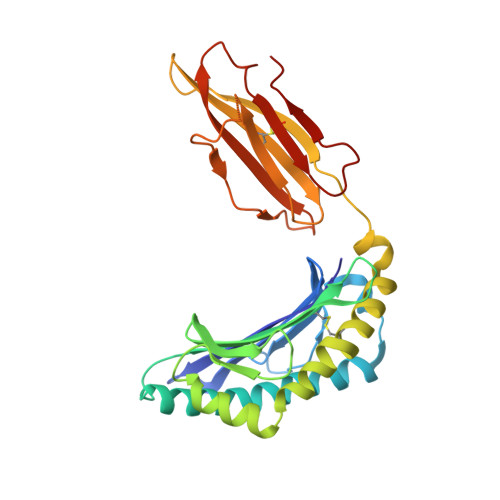
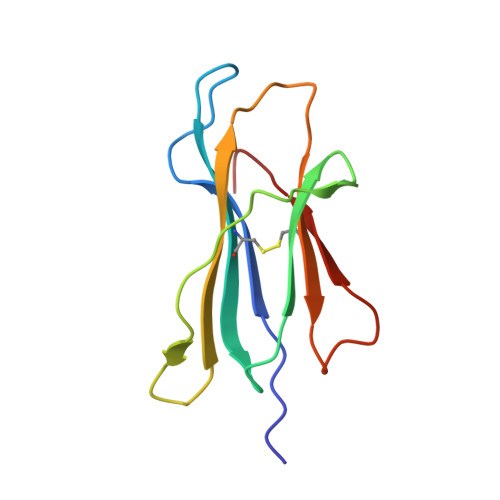
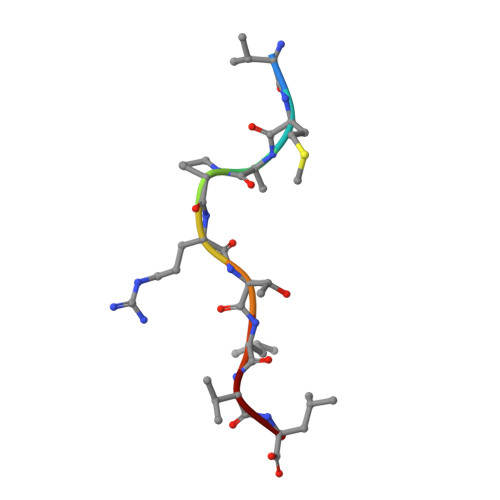
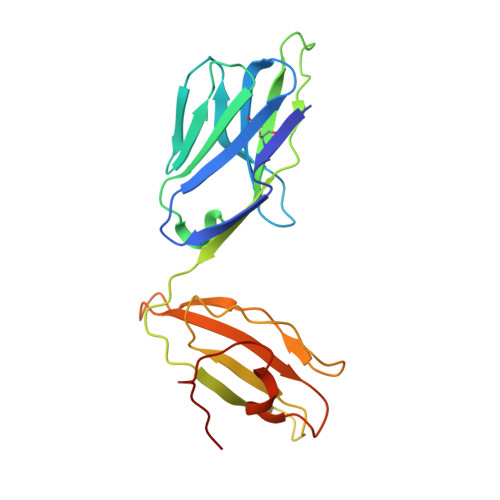
αβ T cell receptors (TCRs) interact with peptides bound to the polymorphic major histocompatibility complex class Ia (MHC-Ia) and class II (MHC-II) molecules as well as the essentially monomorphic MHC class Ib (MHC-Ib) molecules. Although there is a large amount of information on how TCRs engage with MHC-Ia and MHC-II, our understanding of TCR/MHC-Ib interactions is very limited. Infection with cytomegalovirus (CMV) can elicit a CD8 + T cell response restricted by the human MHC-Ib molecule human leukocyte antigen (HLA)-E and specific for an epitope from UL40 (VMAPRTLIL), which is characterized by biased TRBV14 gene usage. Here we describe an HLA-E-restricted CD8 + T cell able to recognize an allotypic variant of the UL40 peptide with a modification at position 8 (P8) of the peptide (VMAPRTLVL) that uses the TRBV9 gene segment. We report the structures of a TRBV9 + TCR in complex with the HLA-E molecule presenting the two peptides. Our data revealed that the TRBV9 + TCR adopts a different docking mode and molecular footprint atop HLA-E when compared with the TRBV14 + TCR-HLA-E ternary complex. Additionally, despite their differing V gene segment usage and different docking mechanisms, mutational analyses showed that the TCRs shared a conserved energetic footprint on the HLA-E molecule, focused around the peptide-binding groove. Hence, we provide new insights into how monomorphic MHC molecules interact with T cells.
- From the Department of Microbiology and Immunology and Peter Doherty Institute for Infection and Immunity, University of Melbourne, Melbourne 3000, Australia.
Organizational Affiliation: