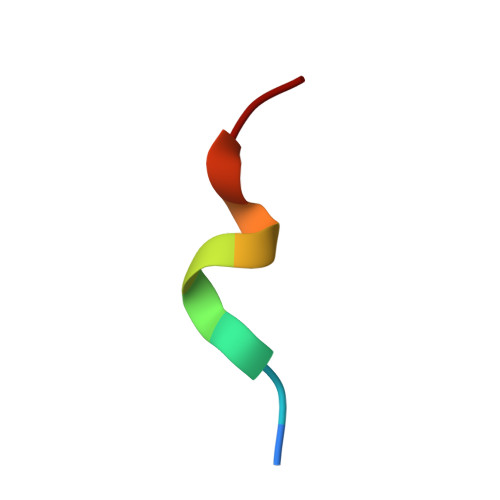A Recurrent Mutation in Anaplastic Lymphoma Kinase with Distinct Neoepitope Conformations.
Toor, J.S., Rao, A.A., McShan, A.C., Yarmarkovich, M., Nerli, S., Yamaguchi, K., Madejska, A.A., Nguyen, S., Tripathi, S., Maris, J.M., Salama, S.R., Haussler, D., Sgourakis, N.G.(2018) Front Immunol 9: 99-99
- PubMed: 29441070 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2018.00099
- Primary Citation Related Structures:
5TXS, 5VZ5, 6AT9 - PubMed Abstract:
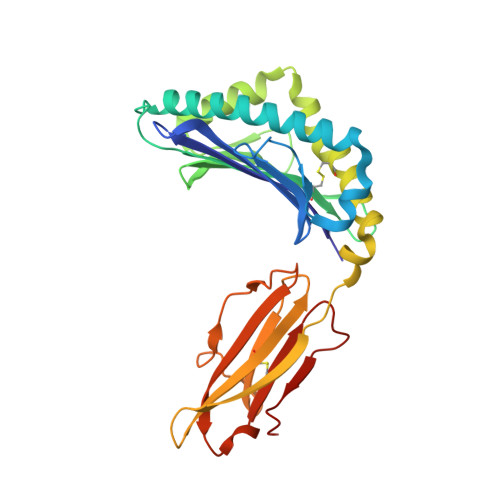
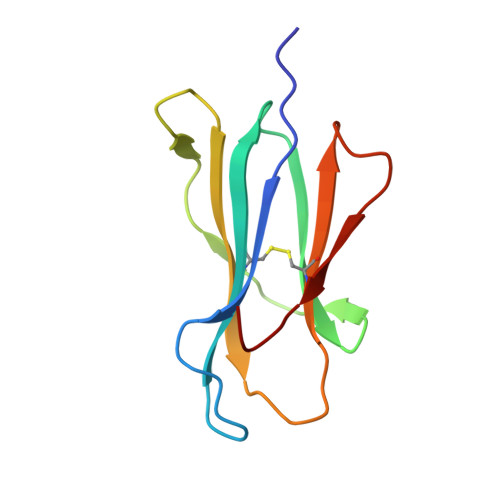
The identification of recurrent human leukocyte antigen (HLA) neoepitopes driving T cell responses against tumors poses a significant bottleneck in the development of approaches for precision cancer therapeutics. Here, we employ a bioinformatics method, Prediction of T Cell Epitopes for Cancer Therapy, to analyze sequencing data from neuroblastoma patients and identify a recurrent anaplastic lymphoma kinase mutation ( ALK R1275Q) that leads to two high affinity neoepitopes when expressed in complex with common HLA alleles. Analysis of the X-ray structures of the two peptides bound to HLA-B*15:01 reveals drastically different conformations with measurable changes in the stability of the protein complexes, while the self-epitope is excluded from binding due to steric hindrance in the MHC groove. To evaluate the range of HLA alleles that could display the ALK neoepitopes, we used structure-based Rosetta comparative modeling calculations, which accurately predict several additional high affinity interactions and compare our results with commonly used prediction tools. Subsequent determination of the X-ray structure of an HLA-A*01:01 bound neoepitope validates atomic features seen in our Rosetta models with respect to key residues relevant for MHC stability and T cell receptor recognition. Finally, MHC tetramer staining of peripheral blood mononuclear cells from HLA-matched donors shows that the two neoepitopes are recognized by CD8+ T cells. This work provides a rational approach toward high-throughput identification and further optimization of putative neoantigen/HLA targets with desired recognition features for cancer immunotherapy.
- Department of Chemistry and Biochemistry, University of California, Santa Cruz, Santa Cruz, CA, United States.
Organizational Affiliation: