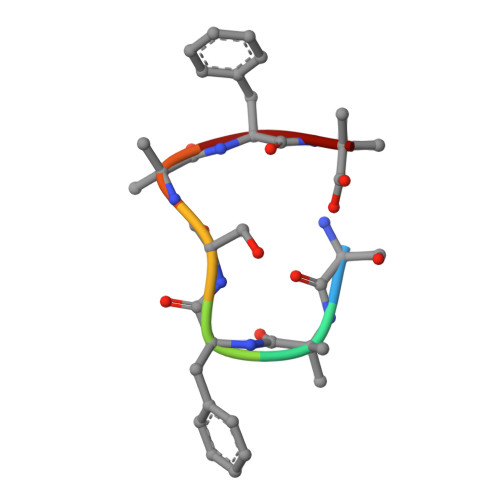
Transition of Metastable Cross-alpha Crystals into Cross-beta Fibrils by beta-Turn Flipping.
Mondal, S., Jacoby, G., Sawaya, M.R., Arnon, Z.A., Adler-Abramovich, L., Rehak, P., Vukovic, L., Shimon, L.J.W., Kral, P., Beck, R., Gazit, E.(2019) J Am Chem Soc 141: 363-369
- PubMed: 30532955 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.8b10289
- Primary Citation Related Structures:
5VSG - PubMed Abstract:
The ensemble of native, folded state was once considered to represent the global energy minimum of a given protein sequence. More recently, the discovery of the cross-β amyloid state revealed that deeper energy minima exist, often associated with pathogenic, fibrillar deposits, when the concentration of proteins reaches a critical value. Fortunately, a sizable energy barrier impedes the conversion from native to pathogenic states. However, little is known about the structure of the related transition state. In addition, there are indications of polymorphism in the amyloidogenic process. Here, we report the first evidence of the conversion of metastable cross-α-helical crystals to thermodynamically stable cross-β-sheet-like fibrils by a de novo designed heptapeptide. Furthermore, for the first time, we demonstrate at atomic resolution that the flip of a peptide plane from a type I to a type II' turn facilitates transformation to cross-β structure and assembly of a dry steric zipper. This study establishes the potential of a peptide turn, a common protein secondary structure, to serve as a principal gatekeeper between a native metastable folded state and the amyloid state.
- Department of Molecular Microbiology and Biotechnology, George S. Wise Faculty of Life Sciences , Tel Aviv University , Tel Aviv 69978 , Israel.
Organizational Affiliation: