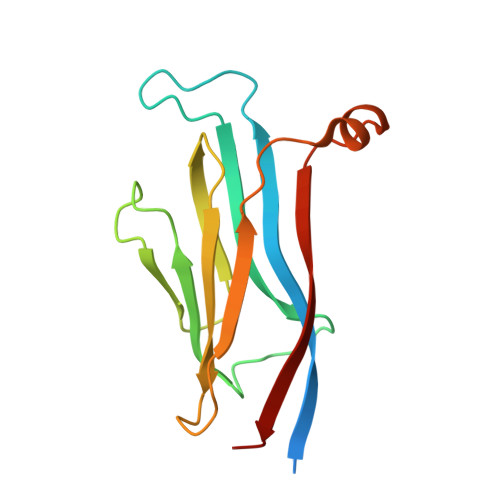
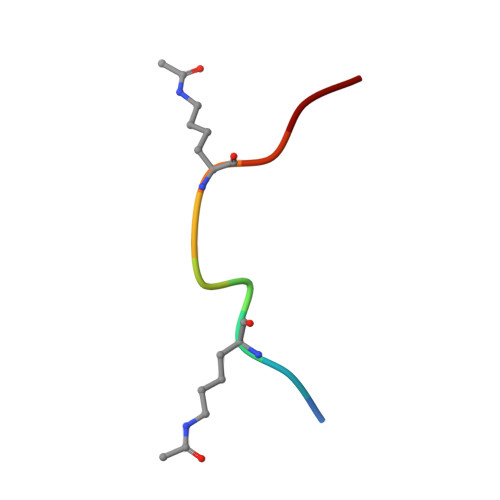
GAS41 Recognizes Diacetylated Histone H3 through a Bivalent Binding Mode.
Cho, H.J., Li, H., Linhares, B.M., Kim, E., Ndoj, J., Miao, H., Grembecka, J., Cierpicki, T.(2018) ACS Chem Biol 13: 2739-2746
- PubMed: 30071723 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.8b00674
- Primary Citation Related Structures:
5VNA, 5VNB - PubMed Abstract:
GAS41 is a chromatin-associated protein that belongs to the YEATS family and is involved in the recognition of acetyl-lysine in histone proteins. A unique feature of GAS41 is the presence of a C-terminal coiled-coil domain, which is responsible for protein dimerization. Here, we characterized the specificity of the GAS41 YEATS domain and found that it preferentially binds to acetylated H3K18 and H3K27 peptides. Interestingly, we found that full-length, dimeric GAS41 binds to diacetylated H3 peptides with an enhanced affinity when compared to those for monoacetylated peptides, through a bivalent binding mode. We determined the crystal structure of the GAS41 YEATS domain with H3K23acK27ac to visualize the molecular basis of diacetylated histone binding. Our results suggest a unique binding mode in which full-length GAS41 is a reader of diacetylated histones.
- Department of Pathology , University of Michigan , Ann Arbor , Michigan 48109 , United States.
Organizational Affiliation: