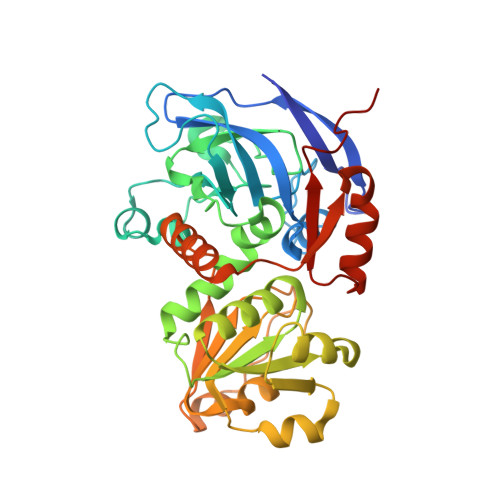
The Enzyme Activity and Substrate Specificity of Two Major Cinnamyl Alcohol Dehydrogenases in Sorghum (Sorghum bicolor), SbCAD2 and SbCAD4.
Jun, S.Y., Walker, A.M., Kim, H., Ralph, J., Vermerris, W., Sattler, S.E., Kang, C.(2017) Plant Physiol 174: 2128-2145
- PubMed: 28606901 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1104/pp.17.00576
- Primary Citation Related Structures:
5VKT - PubMed Abstract:
Cinnamyl alcohol dehydrogenase (CAD) catalyzes the final step in monolignol biosynthesis, reducing sinapaldehyde, coniferaldehyde, and p -coumaraldehyde to their corresponding alcohols in an NADPH-dependent manner. Because of its terminal location in monolignol biosynthesis, the variation in substrate specificity and activity of CAD can result in significant changes in overall composition and amount of lignin. Our in-depth characterization of two major CAD isoforms, SbCAD2 (Brown midrib 6 [bmr6]) and SbCAD4, in lignifying tissues of sorghum ( Sorghum bicolor ), a strategic plant for generating renewable chemicals and fuels, indicates their similarity in both structure and activity to Arabidopsis ( Arabidopsis thaliana ) CAD5 and Populus tremuloides sinapyl alcohol dehydrogenase, respectively. This first crystal structure of a monocot CAD combined with enzyme kinetic data and a catalytic model supported by site-directed mutagenesis allows full comparison with dicot CADs and elucidates the potential signature sequence for their substrate specificity and activity. The L119W/G301F-SbCAD4 double mutant displayed its substrate preference in the order coniferaldehyde > p -coumaraldehyde > sinapaldehyde, with higher catalytic efficiency than that of both wild-type SbCAD4 and SbCAD2. As SbCAD4 is the only major CAD isoform in bmr6 mutants, replacing SbCAD4 with L119W/G301F-SbCAD4 in bmr6 plants could produce a phenotype that is more amenable to biomass processing.
- Department of Chemistry, Washington State University, Pullman, Washington 99164.
Organizational Affiliation: