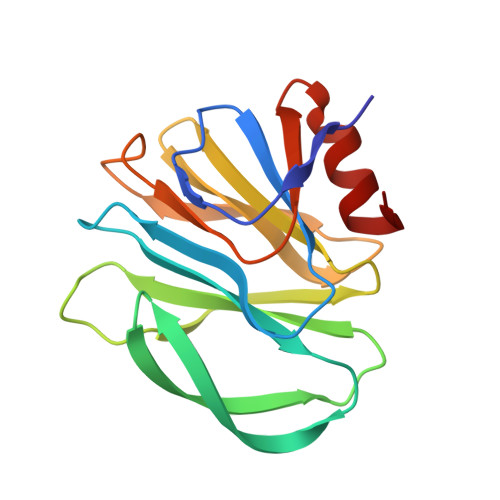
Structural basis of glycan specificity of P[19] VP8*: Implications for rotavirus zoonosis and evolution.
Liu, Y., Xu, S., Woodruff, A.L., Xia, M., Tan, M., Kennedy, M.A., Jiang, X.(2017) PLoS Pathog 13: e1006707-e1006707
- PubMed: 29136651 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1006707
- Primary Citation Related Structures:
5VKI, 5VKS - PubMed Abstract:
Recognition of specific cell surface glycans, mediated by the VP8* domain of the spike protein VP4, is the essential first step in rotavirus (RV) infection. Due to lack of direct structural information of virus-ligand interactions, the molecular basis of ligand-controlled host ranges of the major human RVs (P[8] and P[4]) in P[II] genogroup remains unknown. Here, through characterization of a minor P[II] RV (P[19]) that can infect both animals (pigs) and humans, we made an important advance to fill this knowledge gap by solving the crystal structures of the P[19] VP8* in complex with its ligands. Our data showed that P[19] RVs use a novel binding site that differs from the known ones of other genotypes/genogroups. This binding site is capable of interacting with two types of glycans, the mucin core and type 1 histo-blood group antigens (HBGAs) with a common GlcNAc as the central binding saccharide. The binding site is apparently shared by other P[II] RVs and possibly two genotypes (P[10] and P[12]) in P[I] as shown by their highly conserved GlcNAc-interacting residues. These data provide strong evidence of evolutionary connections among these human and animal RVs, pointing to a common ancestor in P[I] with a possible animal host origin. While the binding properties to GlcNAc-containing saccharides are maintained, changes in binding to additional residues, such as those in the polymorphic type 1 HBGAs may occur in the course of RV evolution, explaining the complex P[II] genogroup that mainly causes diseases in humans but also in some animals.
- Tianjin Key Laboratory of Molecular Nuclear Medicine, Institute of Radiation Medicine, Chinese Academy of Medical Sciences and Peking Union Medical College, Tianjin, China.
Organizational Affiliation: