Structural insights into the mycobacteria transcription initiation complex from analysis of X-ray crystal structures.
Hubin, E.A., Lilic, M., Darst, S.A., Campbell, E.A.(2017) Nat Commun 8: 16072-16072
- PubMed: 28703128 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms16072
- Primary Citation Related Structures:
5VI5, 5VI8 - PubMed Abstract:
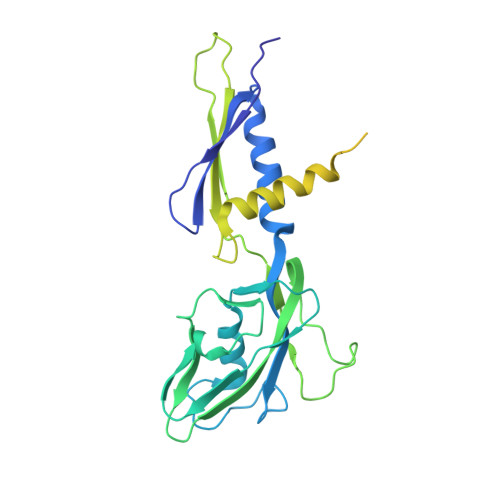
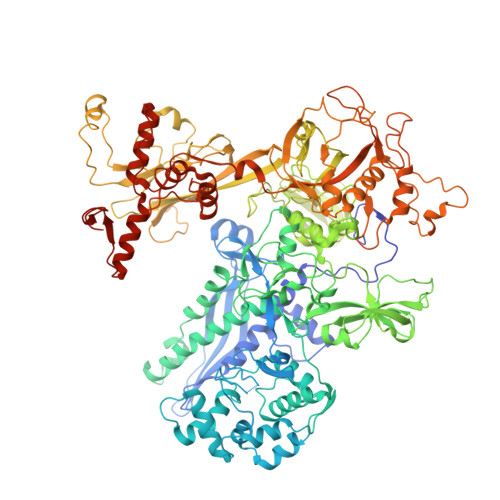
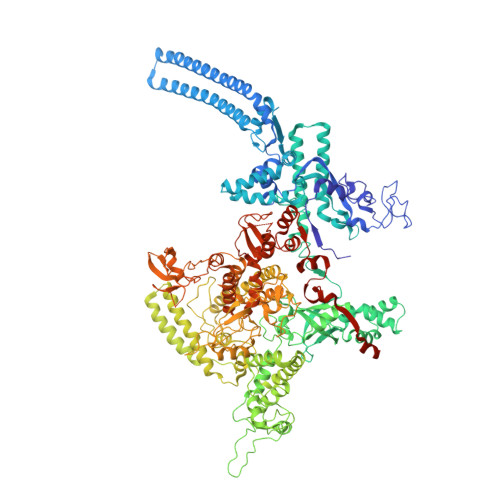
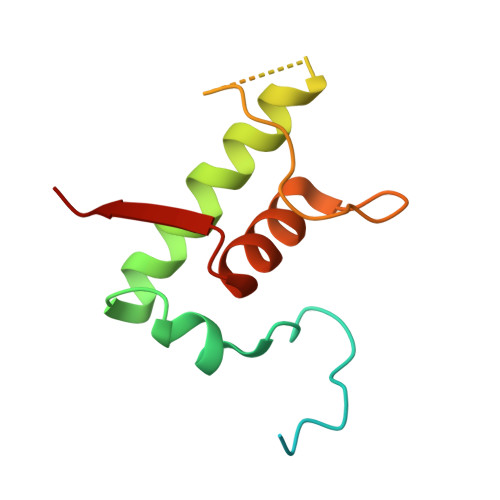
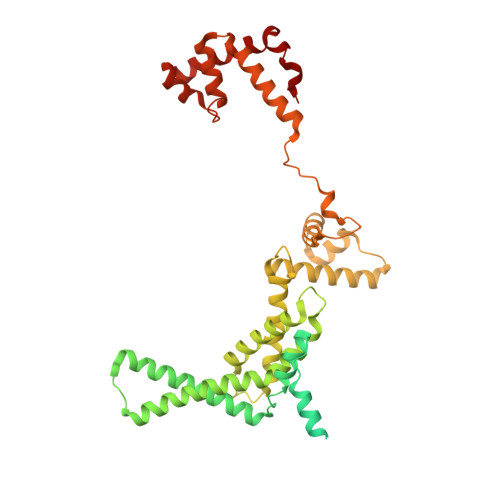
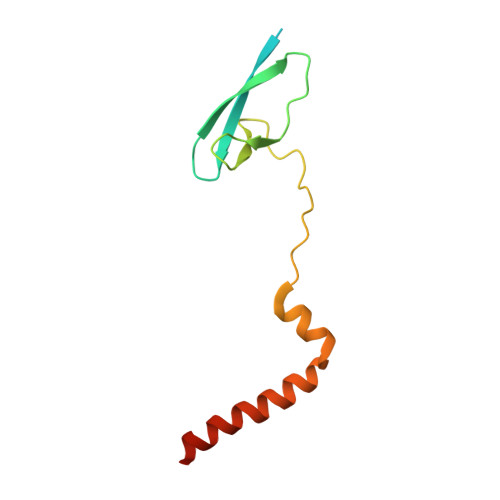
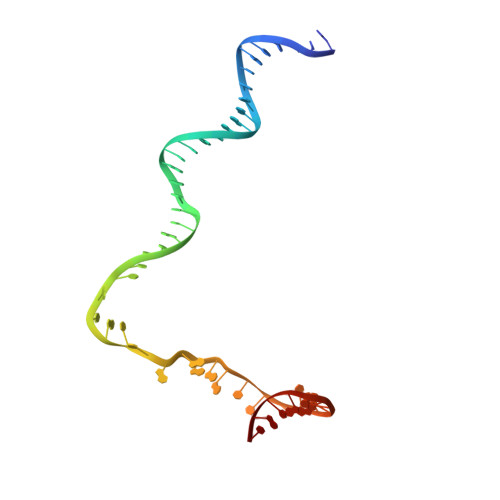
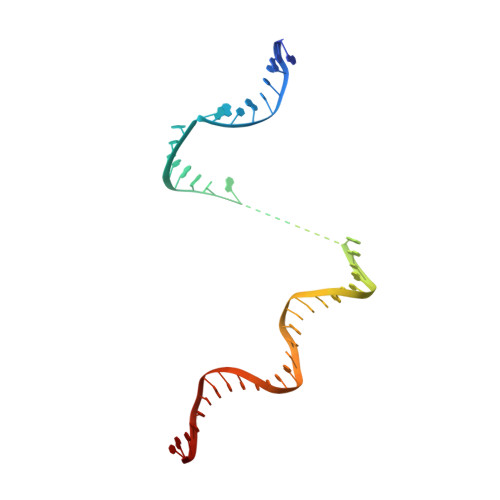
The mycobacteria RNA polymerase (RNAP) is a target for antimicrobials against tuberculosis, motivating structure/function studies. Here we report a 3.2 Å-resolution crystal structure of a Mycobacterium smegmatis (Msm) open promoter complex (RPo), along with structural analysis of the Msm RPo and a previously reported 2.76 Å-resolution crystal structure of an Msm transcription initiation complex with a promoter DNA fragment. We observe the interaction of the Msm RNAP α-subunit C-terminal domain (αCTD) with DNA, and we provide evidence that the αCTD may play a role in Mtb transcription regulation. Our results reveal the structure of an Actinobacteria-unique insert of the RNAP β' subunit. Finally, our analysis reveals the disposition of the N-terminal segment of Msm σ A , which may comprise an intrinsically disordered protein domain unique to mycobacteria. The clade-specific features of the mycobacteria RNAP provide clues to the profound instability of mycobacteria RPo compared with E. coli.
- The Rockefeller University, 1230 York Avenue, New York, New York 10065, USA.
Organizational Affiliation: