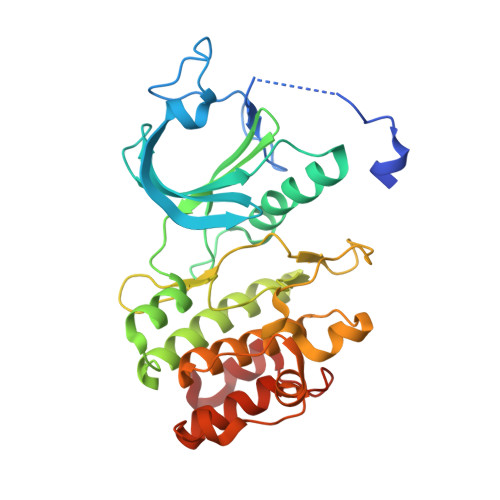
Structural Basis of Wee Kinases Functionality and Inactivation by Diverse Small Molecule Inhibitors.
Zhu, J.Y., Cuellar, R.A., Berndt, N., Lee, H.E., Olesen, S.H., Martin, M.P., Jensen, J.T., Georg, G.I., Schonbrunn, E.(2017) J Med Chem 60: 7863-7875
- PubMed: 28792760 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.7b00996
- Primary Citation Related Structures:
5V5Y, 5VC3, 5VC4, 5VC5, 5VC6, 5VCV, 5VCW, 5VCX, 5VCY, 5VCZ, 5VD0, 5VD1, 5VD2, 5VD3, 5VDK - PubMed Abstract:
Members of the Wee family of kinases negatively regulate the cell cycle via phosphorylation of CDK1 and are considered potential drug targets. Herein, we investigated the structure-function relationship of human Wee1, Wee2, and Myt1 (PKMYT1). Purified recombinant full-length proteins and kinase domain constructs differed substantially in phosphorylation states and catalytic competency, suggesting complex mechanisms of activation. A series of crystal structures reveal unique features that distinguish Wee1 and Wee2 from Myt1 and establish the structural basis of differential inhibition by the widely used Wee1 inhibitor MK-1775. Kinome profiling and cellular studies demonstrate that, in addition to Wee1 and Wee2, MK-1775 is an equally potent inhibitor of the polo-like kinase PLK1. Several previously unrecognized inhibitors of Wee kinases were discovered and characterized. Combined, the data provide a comprehensive view on the catalytic and structural properties of Wee kinases and a framework for the rational design of novel inhibitors thereof.
- Drug Discovery Department, Moffitt Cancer Center , Tampa, Florida 33612, United States.
Organizational Affiliation: