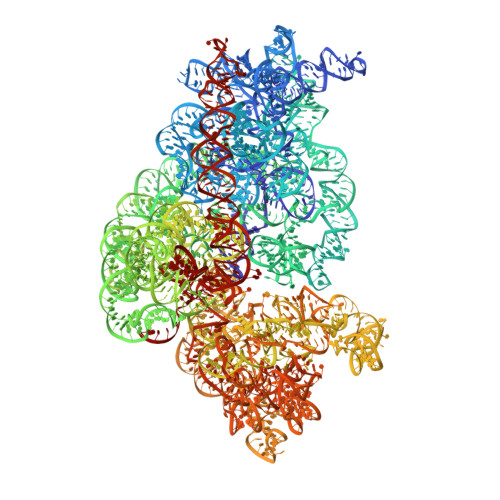
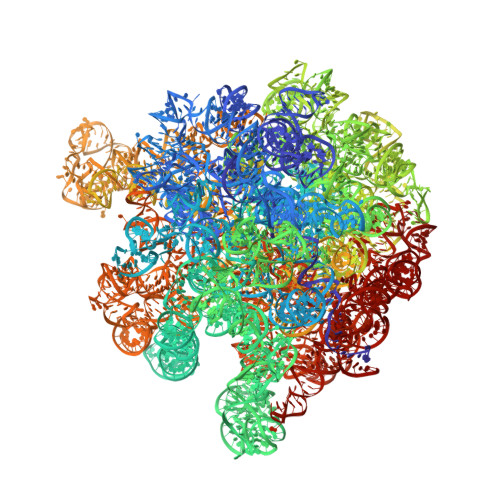
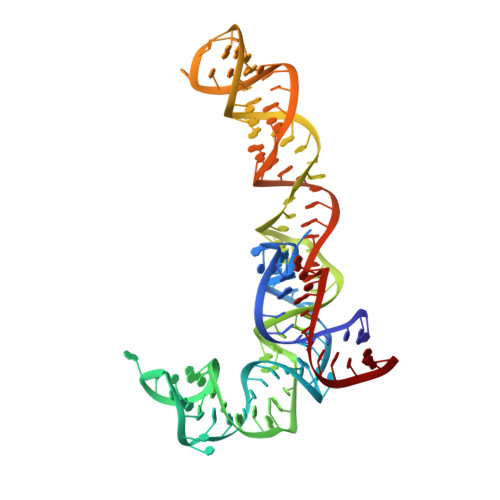
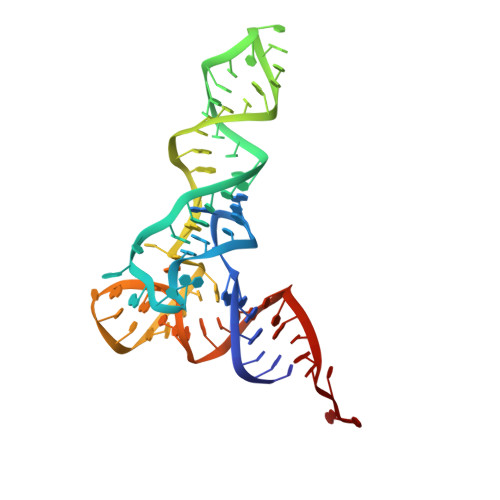
Ensemble cryo-EM elucidates the mechanism of translation fidelity
Loveland, A.B., Demo, G., Grigorieff, N., Korostelev, A.A.(2017) Nature 546: 113-117
- PubMed: 28538735 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature22397
- Primary Citation Related Structures:
5UYK, 5UYL, 5UYM, 5UYN, 5UYP, 5UYQ - PubMed Abstract:
Gene translation depends on accurate decoding of mRNA, the structural mechanism of which remains poorly understood. Ribosomes decode mRNA codons by selecting cognate aminoacyl-tRNAs delivered by elongation factor Tu (EF-Tu). Here we present high-resolution structural ensembles of ribosomes with cognate or near-cognate aminoacyl-tRNAs delivered by EF-Tu. Both cognate and near-cognate tRNA anticodons explore the aminoacyl-tRNA-binding site (A site) of an open 30S subunit, while inactive EF-Tu is separated from the 50S subunit. A transient conformation of decoding-centre nucleotide G530 stabilizes the cognate codon-anticodon helix, initiating step-wise 'latching' of the decoding centre. The resulting closure of the 30S subunit docks EF-Tu at the sarcin-ricin loop of the 50S subunit, activating EF-Tu for GTP hydrolysis and enabling accommodation of the aminoacyl-tRNA. By contrast, near-cognate complexes fail to induce the G530 latch, thus favouring open 30S pre-accommodation intermediates with inactive EF-Tu. This work reveals long-sought structural differences between the pre-accommodation of cognate and near-cognate tRNAs that elucidate the mechanism of accurate decoding.
- RNA Therapeutics Institute, Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, 368 Plantation Street, Worcester, Massachusetts 01605, USA.
Organizational Affiliation: