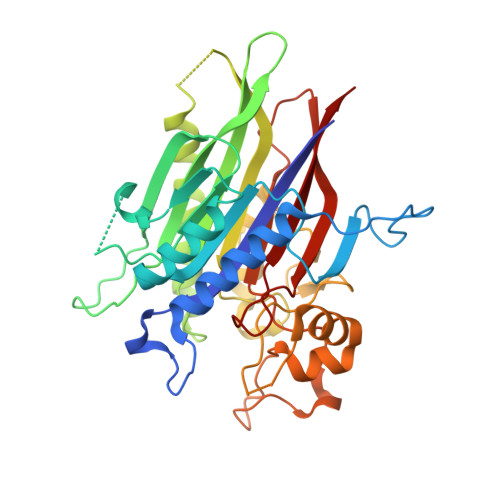
Structure of human nSMase2 reveals an interdomain allosteric activation mechanism for ceramide generation.
Airola, M.V., Shanbhogue, P., Shamseddine, A.A., Guja, K.E., Senkal, C.E., Maini, R., Bartke, N., Wu, B.X., Obeid, L.M., Garcia-Diaz, M., Hannun, Y.A.(2017) Proc Natl Acad Sci U S A 114: E5549-E5558
- PubMed: 28652336 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1705134114
- Primary Citation Related Structures:
5UVG - PubMed Abstract:
Neutral sphingomyelinase 2 (nSMase2, product of the SMPD3 gene) is a key enzyme for ceramide generation that is involved in regulating cellular stress responses and exosome-mediated intercellular communication. nSMase2 is activated by diverse stimuli, including the anionic phospholipid phosphatidylserine. Phosphatidylserine binds to an integral-membrane N-terminal domain (NTD); however, how the NTD activates the C-terminal catalytic domain is unclear. Here, we identify the complete catalytic domain of nSMase2, which was misannotated because of a large insertion. We find the soluble catalytic domain interacts directly with the membrane-associated NTD, which serves as both a membrane anchor and an allosteric activator. The juxtamembrane region, which links the NTD and the catalytic domain, is necessary and sufficient for activation. Furthermore, we provide a mechanistic basis for this phenomenon using the crystal structure of the human nSMase2 catalytic domain determined at 1.85-Å resolution. The structure reveals a DNase-I-type fold with a hydrophobic track leading to the active site that is blocked by an evolutionarily conserved motif which we term the "DK switch." Structural analysis of nSMase2 and the extended N-SMase family shows that the DK switch can adopt different conformations to reposition a universally conserved Asp (D) residue involved in catalysis. Mutation of this Asp residue in nSMase2 disrupts catalysis, allosteric activation, stimulation by phosphatidylserine, and pharmacological inhibition by the lipid-competitive inhibitor GW4869. Taken together, these results demonstrate that the DK switch regulates ceramide generation by nSMase2 and is governed by an allosteric interdomain interaction at the membrane interface.
- Stony Brook University Cancer Center, Stony Brook, NY 11794.
Organizational Affiliation: