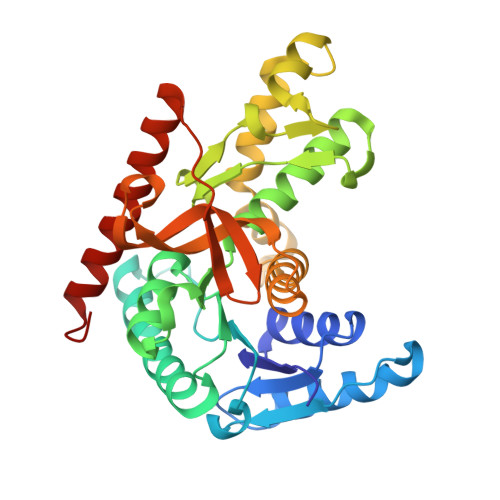
Conformational changes on substrate binding revealed by structures of Methylobacterium extorquens malate dehydrogenase.
Gonzalez, J.M., Marti-Arbona, R., Chen, J.C.H., Broom-Peltz, B., Unkefer, C.J.(2018) Acta Crystallogr F Struct Biol Commun 74: 610-616
- PubMed: 30279311 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18011809
- Primary Citation Related Structures:
5UJK, 5ULV - PubMed Abstract:
Three high-resolution X-ray crystal structures of malate dehydrogenase (MDH; EC 1.1.1.37) from the methylotroph Methylobacterium extorquens AM1 are presented. By comparing the structures of apo MDH, a binary complex of MDH and NAD + , and a ternary complex of MDH and oxaloacetate with ADP-ribose occupying the pyridine nucleotide-binding site, conformational changes associated with the formation of the catalytic complex were characterized. While the substrate-binding site is accessible in the enzyme resting state or NAD + -bound forms, the substrate-bound form exhibits a closed conformation. This conformational change involves the transition of an α-helix to a 3 10 -helix, which causes the adjacent loop to close the active site following coenzyme and substrate binding. In the ternary complex, His284 forms a hydrogen bond to the C2 carbonyl of oxaloacetate, placing it in a position to donate a proton in the formation of (2S)-malate.
- Instituto de Bionanotecnología del NOA, Consejo Nacional de Investigaciones Científicas y Técnicas, Universidad Nacional de Santiago del Estero, G4206XCP Santiago del Estero, Argentina.
Organizational Affiliation: