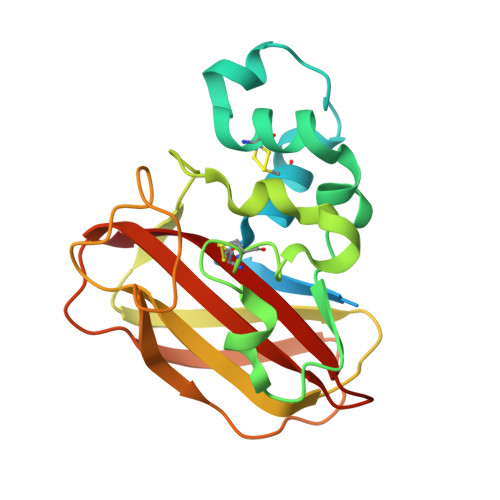
Structure of a Thermobifida fusca lytic polysaccharide monooxygenase and mutagenesis of key residues.
Kruer-Zerhusen, N., Alahuhta, M., Lunin, V.V., Himmel, M.E., Bomble, Y.J., Wilson, D.B.(2017) Biotechnol Biofuels 10: 243-243
- PubMed: 29213309
- DOI: https://doi.org/10.1186/s13068-017-0925-7
- Primary Citation Related Structures:
5UIZ - PubMed Abstract:
Auxiliary activity (AA) enzymes are produced by numerous bacterial and fungal species to assist in the degradation of biomass. These enzymes are abundant but have yet to be fully characterized. Here, we report the X-ray structure of Thermobifida fusca AA10A (TfAA10A), investigate mutational characterization of key surface residues near its active site, and explore the importance of the various domains of Thermobifida fusca AA10B (TfAA10B). The structure of TfAA10A is similar to other bacterial LPMOs (lytic polysaccharide monooxygenases), including signs of photo-reduction and a distorted active site, with mixed features showing both type I and II copper coordination. The point mutation experiments of TfAA10A show that Trp82 and Asn83 are needed for binding, but only Trp82 affects activity. The TfAA10B domain truncation mutants reveal that CBM2 is crucial for the binding of substrate, but that the X1 module does not affect binding or activity. In TfAA10A, Trp82 and Asn83 are needed for binding, but only Trp82 affects activity. The TfAA10B domain truncation mutants reveal that CBM2 is crucial for substrate binding, but that the X1 module does not affect binding or activity. The structure of TfAA10A is similar to other bacterial lytic polysaccharide monooxygenases with mixed features showing both type I and II copper coordination. The role of LPMOs and the variability of abundance in genomes are not fully explored. LPMOs likely perform initial attacks into crystalline cellulose to allow larger processive cellulases to bind and attack, but the precise nature of their synergistic behavior remains to be definitively characterized.
- Department of Molecular Biology and Genetics, Cornell University, Ithaca, NY USA.
Organizational Affiliation: