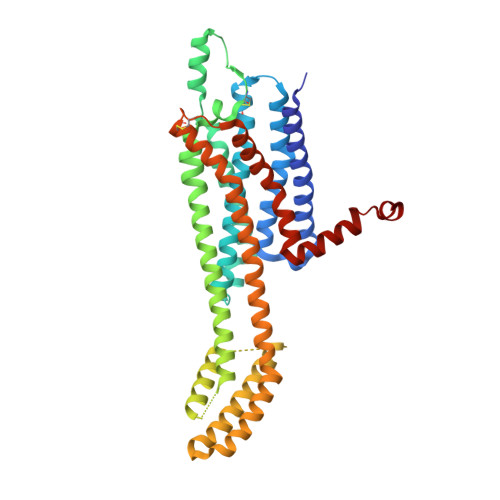
Structure of the Adenosine A1 Receptor Reveals the Basis for Subtype Selectivity.
Glukhova, A., Thal, D.M., Nguyen, A.T., Vecchio, E.A., Jorg, M., Scammells, P.J., May, L.T., Sexton, P.M., Christopoulos, A.(2017) Cell 168: 867-877.e13
- PubMed: 28235198 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2017.01.042
- Primary Citation Related Structures:
5UEN - PubMed Abstract:
The adenosine A 1 receptor (A 1 -AR) is a G-protein-coupled receptor that plays a vital role in cardiac, renal, and neuronal processes but remains poorly targeted by current drugs. We determined a 3.2 Å crystal structure of the A 1 -AR bound to the selective covalent antagonist, DU172, and identified striking differences to the previously solved adenosine A 2A receptor (A 2A -AR) structure. Mutational and computational analysis of A 1 -AR revealed a distinct conformation of the second extracellular loop and a wider extracellular cavity with a secondary binding pocket that can accommodate orthosteric and allosteric ligands. We propose that conformational differences in these regions, rather than amino-acid divergence, underlie drug selectivity between these adenosine receptor subtypes. Our findings provide a molecular basis for AR subtype selectivity with implications for understanding the mechanisms governing allosteric modulation of these receptors, allowing the design of more selective agents for the treatment of ischemia-reperfusion injury, renal pathologies, and neuropathic pain.
- Drug Discovery Biology, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville, Victoria 3052, Australia.
Organizational Affiliation: