Drugs and drug-like molecules can modulate the function of mucosal-associated invariant T cells.
Keller, A.N., Eckle, S.B., Xu, W., Liu, L., Hughes, V.A., Mak, J.Y., Meehan, B.S., Pediongco, T., Birkinshaw, R.W., Chen, Z., Wang, H., D'Souza, C., Kjer-Nielsen, L., Gherardin, N.A., Godfrey, D.I., Kostenko, L., Corbett, A.J., Purcell, A.W., Fairlie, D.P., McCluskey, J., Rossjohn, J.(2017) Nat Immunol 18: 402-411
- PubMed: 28166217 Search on PubMed
- DOI: https://doi.org/10.1038/ni.3679
- Primary Citation Related Structures:
5U16, 5U17, 5U1R, 5U2V, 5U6Q - PubMed Abstract:
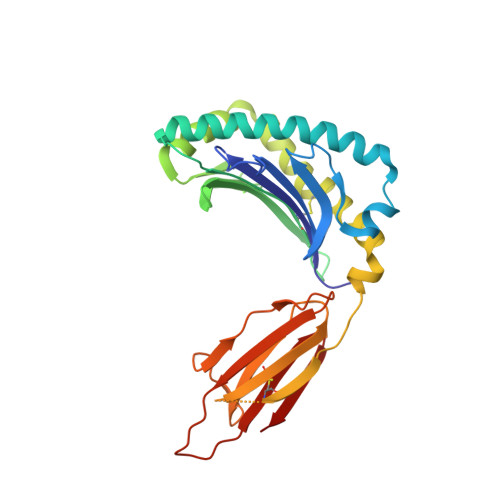
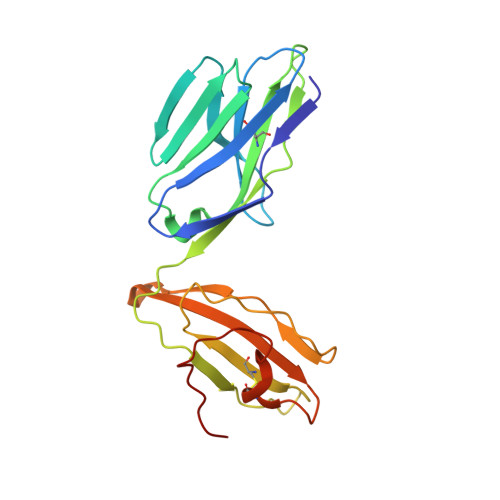
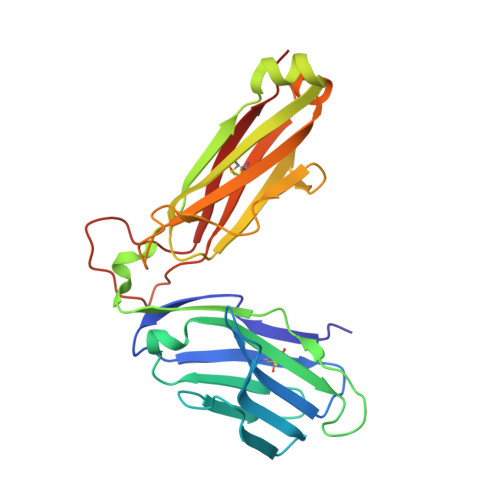
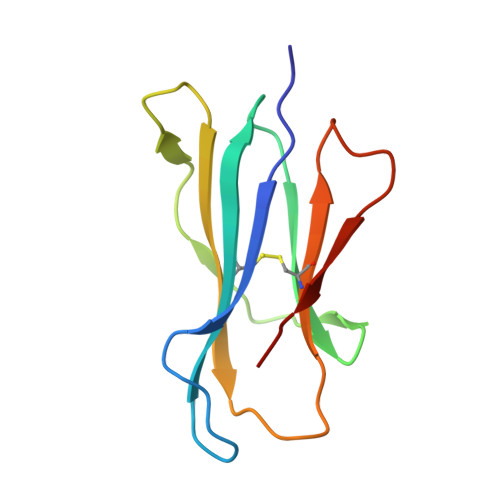
The major-histocompatibility-complex-(MHC)-class-I-related molecule MR1 can present activating and non-activating vitamin-B-based ligands to mucosal-associated invariant T cells (MAIT cells). Whether MR1 binds other ligands is unknown. Here we identified a range of small organic molecules, drugs, drug metabolites and drug-like molecules, including salicylates and diclofenac, as MR1-binding ligands. Some of these ligands inhibited MAIT cells ex vivo and in vivo, while others, including diclofenac metabolites, were agonists. Crystal structures of a T cell antigen receptor (TCR) from a MAIT cell in complex with MR1 bound to the non-stimulatory and stimulatory compounds showed distinct ligand orientations and contacts within MR1, which highlighted the versatility of the MR1 binding pocket. The findings demonstrated that MR1 was able to capture chemically diverse structures, spanning mono- and bicyclic compounds, that either inhibited or activated MAIT cells. This indicated that drugs and drug-like molecules can modulate MAIT cell function in mammals.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Australia.
Organizational Affiliation: