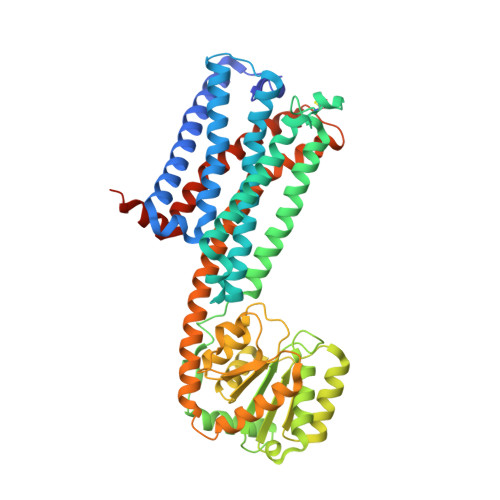
High-resolution crystal structure of the human CB1 cannabinoid receptor.
Shao, Z., Yin, J., Chapman, K., Grzemska, M., Clark, L., Wang, J., Rosenbaum, D.M.(2016) Nature
- PubMed: 27851727 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature20613
- Primary Citation Related Structures:
5U09 - PubMed Abstract:
The human cannabinoid G-protein-coupled receptors (GPCRs) CB1 and CB2 mediate the functional responses to the endocannabinoids anandamide and 2-arachidonyl glycerol (2-AG) and to the widely consumed plant phytocannabinoid Δ 9 -tetrahydrocannabinol (THC). The cannabinoid receptors have been the targets of intensive drug discovery efforts, because modulation of these receptors has therapeutic potential to control pain, epilepsy, obesity, and other disorders. Although much progress in understanding the biophysical properties of GPCRs has recently been made, investigations of the molecular mechanisms of the cannabinoids and their receptors have lacked high-resolution structural data. Here we report the use of GPCR engineering and lipidic cubic phase crystallization to determine the structure of the human CB1 receptor bound to the inhibitor taranabant at 2.6-Å resolution. We found that the extracellular surface of CB1, including the highly conserved membrane-proximal N-terminal region, is distinct from those of other lipid-activated GPCRs, forming a critical part of the ligand-binding pocket. Docking studies further demonstrate how this same pocket may accommodate the cannabinoid agonist THC. Our CB1 structure provides an atomic framework for studying cannabinoid receptor function and will aid the design and optimization of therapeutic modulators of the endocannabinoid system.
- Department of Biophysics, The University of Texas Southwestern Medical Center, Dallas, Texas 75390, USA.
Organizational Affiliation: