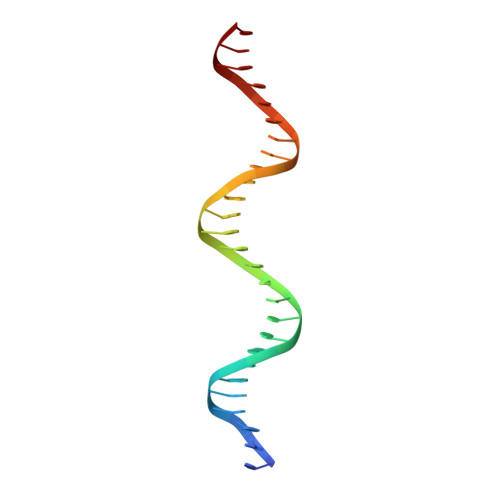
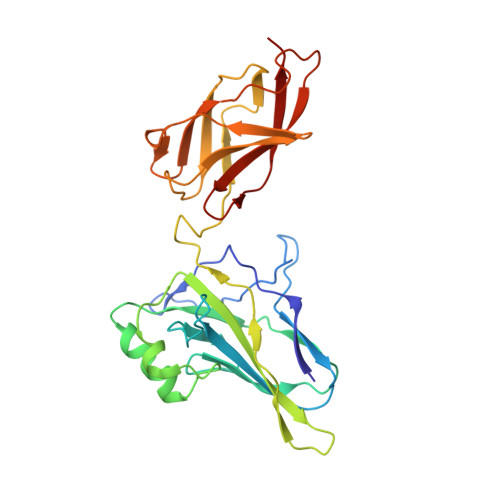
DNA-binding affinity and transcriptional activity of the RelA homodimer of nuclear factor kappa B are not correlated.
Mulero, M.C., Huang, D.B., Nguyen, H.T., Wang, V.Y., Li, Y., Biswas, T., Ghosh, G.(2017) J Biological Chem 292: 18821-18830
- PubMed: 28935669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.813980
- Primary Citation Related Structures:
5U01 - PubMed Abstract:
The nuclear factor κB (NF-κB) transcription factor family regulates genes involved in cell proliferation and inflammation. The promoters of these genes often contain NF-κB-binding sites (κB sites) arranged in tandem. How NF-κB activates transcription through these multiple sites is incompletely understood. We report here an X-ray crystal structure of homodimers comprising the RelA DNA-binding domain containing the Rel homology region (RHR) in NF-κB bound to an E-selectin promoter fragment with tandem κB sites. This structure revealed that two dimers bind asymmetrically to the symmetrically arranged κB sites at which multiple cognate contacts between one dimer to the corresponding DNA are broken. Because simultaneous RelA-RHR dimer binding to tandem sites in solution was anti-cooperative, we inferred that asymmetric RelA-RHR binding with fewer contacts likely indicates a dissociative binding mode. We found that both κB sites are essential for reporter gene activation by full-length RelA homodimer, suggesting that dimers facilitate DNA binding to each other even though their stable co-occupation is not promoted. Promoter variants with altered spacing and orientation of tandem κB sites displayed unexpected reporter activities that were not explained by the solution-binding pattern of RelA-RHR. Remarkably, full-length RelA bound all DNAs with a weaker affinity and specificity. Moreover, the transactivation domain played a negative role in DNA binding. These observations suggest that other nuclear factors influence full-length RelA binding to DNA by neutralizing the transactivation domain negative effect. We propose that DNA binding by NF-κB dimers is highly complex and modulated by facilitated association-dissociation processes.
- From the Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla, California 92093 and.
Organizational Affiliation: