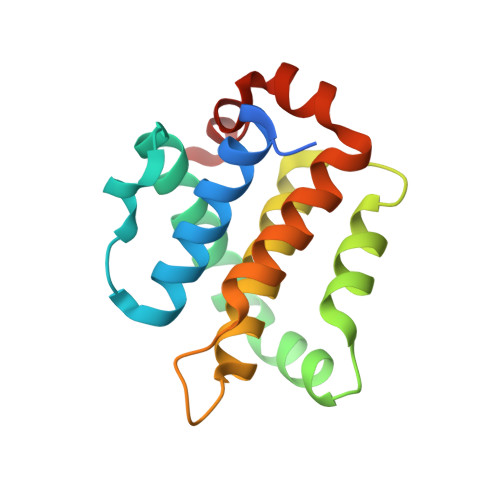
Structural basis of apoptosis inhibition by the fowlpox virus protein FPV039.
Anasir, M.I., Caria, S., Skinner, M.A., Kvansakul, M.(2017) J Biological Chem 292: 9010-9021
- PubMed: 28411240 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.768879
- Primary Citation Related Structures:
5TZP, 5TZQ - PubMed Abstract:
Programmed cell death or apoptosis of infected host cells is an important defense mechanism in response to viral infections. This process is regulated by proapoptotic and prosurvival members of the B-cell lymphoma 2 (Bcl-2) protein family. To counter premature death of a virus-infected cell, poxviruses use a range of different molecular strategies including the mimicry of prosurvival Bcl-2 proteins. One such viral prosurvival protein is the fowlpox virus protein FPV039, which is a potent apoptosis inhibitor, but the precise molecular mechanism by which FPV039 inhibits apoptosis is unknown. To understand how fowlpox virus inhibits apoptosis, we examined FPV039 using isothermal titration calorimetry, small-angle X-ray scattering, and X-ray crystallography. Here, we report that the fowlpox virus prosurvival protein FPV039 promiscuously binds to cellular proapoptotic Bcl-2 and engages all major proapoptotic Bcl-2 proteins. Unlike other identified viral Bcl-2 proteins to date, FPV039 engaged with cellular proapoptotic Bcl-2 with affinities comparable with those of Bcl-2's endogenous cellular counterparts. Structural studies revealed that FPV039 adopts the conserved Bcl-2 fold observed in cellular prosurvival Bcl-2 proteins and closely mimics the structure of the prosurvival Bcl-2 family protein Mcl-1. Our findings suggest that FPV039 is a pan-Bcl-2 protein inhibitor that can engage all host BH3-only proteins, as well as Bcl-2-associated X, apoptosis regulator (Bax) and Bcl-2 antagonist/killer (Bak) proteins to inhibit premature apoptosis of an infected host cell. This work therefore provides a mechanistic platform to better understand FPV039-mediated apoptosis inhibition.
- From the Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, Victoria 3086, Australia and.
Organizational Affiliation: