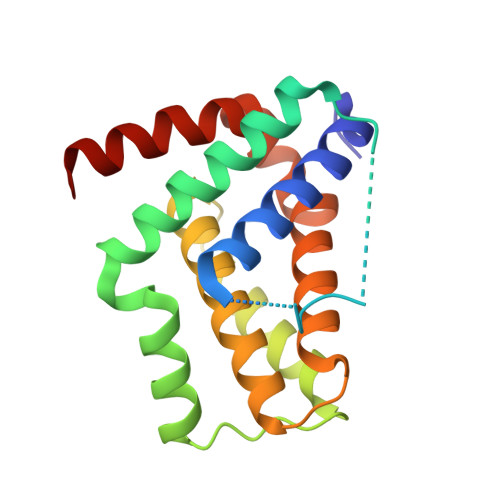
Structural insight into an evolutionarily ancient programmed cell death regulator - the crystal structure of marine sponge BHP2 bound to LB-Bak-2.
Caria, S., Hinds, M.G., Kvansakul, M.(2017) Cell Death Dis 8: e2543-e2543
- PubMed: 28079890 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cddis.2016.469
- Primary Citation Related Structures:
5TWA - PubMed Abstract:
Sponges of the porifera family harbor some of the evolutionary most ancient orthologs of the B-cell lymphoma-2 (Bcl-2) family, a protein family critical to regulation of apoptosis. The genome of the sponge Geodia cydonium contains the putative pro-survival Bcl-2 homolog BHP2, which protects sponge tissue as well as mammalian Hek-293 and NIH-3T3 cells against diverse apoptotic stimuli. The Lake Baikal demosponge Lubomirskia baicalensis has been shown to encode both putative pro-survival Bcl-2 (LB-Bcl-2) and pro-apoptotic Bcl-2 members (LB-Bak-2), which have been implied in axis formation (branches) in L. baicalensis. However, the molecular mechanism of action of sponge-encoded orthologs of Bcl-2 remains to be clarified. Here, we report that the pro-survival Bcl-2 ortholog BHP2 from G. cydonium is able to bind the BH3 motif of a pro-apoptotic Bcl-2 protein, LB-Bak-2 of the sponge L. baicalensis. Furthermore, we determined the crystal structure of BHP2 bound to LB-Bak-2, which revealed that using a binding groove conserved across all pro-survival Bcl-2 proteins, BHP2 binds multi-motif Bax-like proteins through their BH3-binding regions. However, BHP2 discriminates against BH3-only bearing proteins by blocking access to a hydrophobic pocket that is critical for BH3 motif binding in pro-survival Bcl-2 proteins from higher organisms. This differential binding mode is reflected in a structure-based phylogenetic comparison of BHP2 with other Bcl-2 family members, which revealed that BHP2 does not cluster with either Bcl-2 members of higher organisms or pathogen-encoded homologs, and assumes a discrete position. Our findings suggest that the molecular machinery and mechanisms for executing Bcl-2-mediated apoptosis as observed in mammals are evolutionary ancient, with early regulation of apoptotic machineries closely resembling their modern counterparts in mammals rather than Caenorhabditis elegans or drosophila.
- Department of Biochemistry & Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne,Victoria 3086, Australia.
Organizational Affiliation: