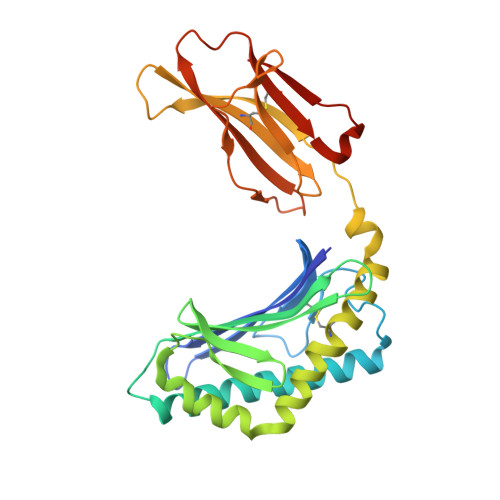
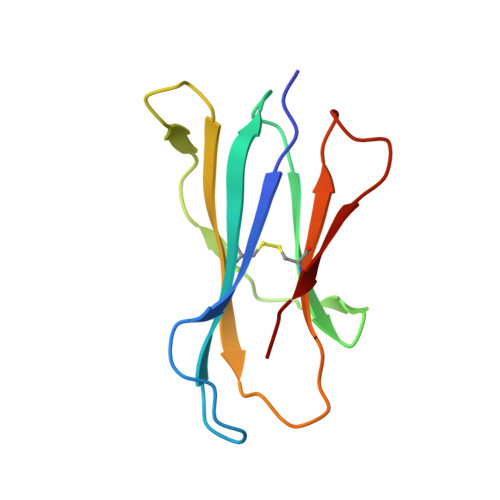
Galactosylsphingamides: new alpha-GalCer analogues to probe the F'-pocket of CD1d.
Guillaume, J., Wang, J., Janssens, J., Remesh, S.G., Risseeuw, M.D.P., Decruy, T., Froeyen, M., Elewaut, D., Zajonc, D.M., Calenbergh, S.V.(2017) Sci Rep 7: 4276-4276
- PubMed: 28655912 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-04461-7
- Primary Citation Related Structures:
5TW2, 5TW5 - PubMed Abstract:
Invariant Natural Killer T-cells (iNKT-cells) are an attractive target for immune response modulation, as upon CD1d-mediated stimulation with KRN7000, a synthetic α-galactosylceramide, they produce a vast amount of cytokines. Here we present a synthesis that allows swift modification of the phytosphingosine side chain by amidation of an advanced methyl ester precursor. The resulting KRN7000 derivatives, termed α-galactosylsphingamides, were evaluated for their capacity to stimulate iNKT-cells. While introduction of the amide-motif in the phytosphingosine chain is tolerated for CD1d binding and TCR recognition, the studied α-galactosylsphingamides showed compromised antigenic properties.
- Laboratory for Medicinal Chemistry (FFW), Faculty of Pharmaceutical Sciences, UGent, Ottergemsesteenweg 460, B-9000, Ghent, Belgium.
Organizational Affiliation: