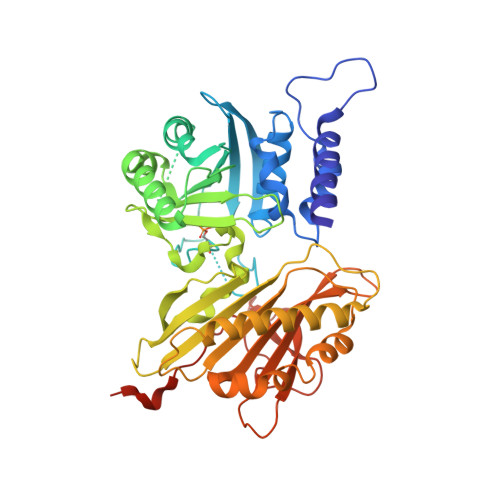
A unified molecular mechanism for the regulation of acetyl-CoA carboxylase by phosphorylation.
Wei, J., Zhang, Y., Yu, T.Y., Sadre-Bazzaz, K., Rudolph, M.J., Amodeo, G.A., Symington, L.S., Walz, T., Tong, L.(2016) Cell Discov 2: 16044-16044
- PubMed: 27990296 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/celldisc.2016.44
- Primary Citation Related Structures:
5TRC - PubMed Abstract:
Acetyl-CoA carboxylases (ACCs) are crucial metabolic enzymes and attractive targets for drug discovery. Eukaryotic acetyl-CoA carboxylases are 250 kDa single-chain, multi-domain enzymes and function as dimers and higher oligomers. Their catalytic activity is tightly regulated by phosphorylation and other means. Here we show that yeast ACC is directly phosphorylated by the protein kinase SNF1 at residue Ser1157, which potently inhibits the enzyme. Crystal structure of three ACC central domains (AC3-AC5) shows that the phosphorylated Ser1157 is recognized by Arg1173, Arg1260, Tyr1113 and Ser1159. The R1173A/R1260A double mutant is insensitive to SNF1, confirming that this binding site is crucial for regulation. Electron microscopic studies reveal dramatic conformational changes in the holoenzyme upon phosphorylation, likely owing to the dissociation of the biotin carboxylase domain dimer. The observations support a unified molecular mechanism for the regulation of ACC by phosphorylation as well as by the natural product soraphen A, a potent inhibitor of eukaryotic ACC. These molecular insights enhance our understanding of acetyl-CoA carboxylase regulation and provide a basis for drug discovery.
- Department of Biological Sciences, Columbia University , New York, NY, USA.
Organizational Affiliation: