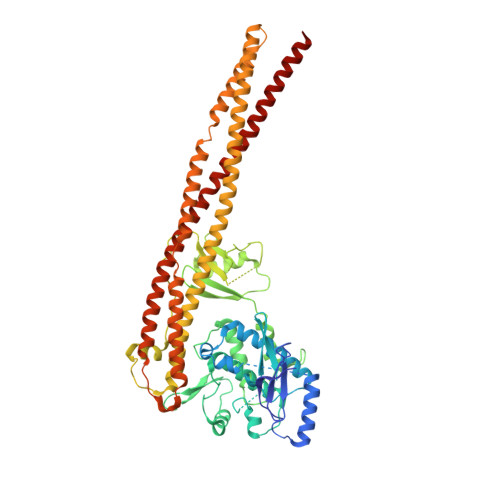
Structural Basis for the Activation of IKK1/ alpha.
Polley, S., Passos, D.O., Huang, D.B., Mulero, M.C., Mazumder, A., Biswas, T., Verma, I.M., Lyumkis, D., Ghosh, G.(2016) Cell Rep 17: 1907-1914
- PubMed: 27851956 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2016.10.067
- Primary Citation Related Structures:
5EBZ, 5TQW, 5TQX, 5TQY - PubMed Abstract:
Distinct signaling pathways activate the NF-κB family of transcription factors. The canonical NF-κB-signaling pathway is mediated by IκB kinase 2/β (IKK2/β), while the non-canonical pathway depends on IKK1/α. The structural and biochemical bases for distinct signaling by these otherwise highly similar IKKs are unclear. We report single-particle cryoelectron microscopy (cryo-EM) and X-ray crystal structures of human IKK1 in dimeric (∼150 kDa) and hexameric (∼450 kDa) forms. The hexamer, which is the representative form in the crystal but comprises only ∼2% of the particles in solution by cryo-EM, is a trimer of IKK1 dimers. While IKK1 hexamers are not detectable in cells, the surface that supports hexamer formation is critical for IKK1-dependent cellular processing of p100 to p52, the hallmark of non-canonical NF-κB signaling. Comparison of this surface to that in IKK2 indicates significant divergence, and it suggests a fundamental role for this surface in signaling by these kinases through distinct pathways.
- Department of Chemistry & Biochemistry, University of California San Diego, La Jolla, CA 92093, USA; Laboratory of Genetics and Helmsley Center for Genomic Medicine, The Salk Institute for Biological Studies, La Jolla, CA 92037, USA.
Organizational Affiliation: