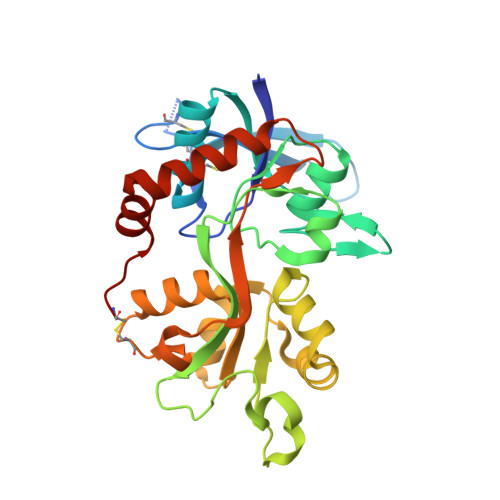
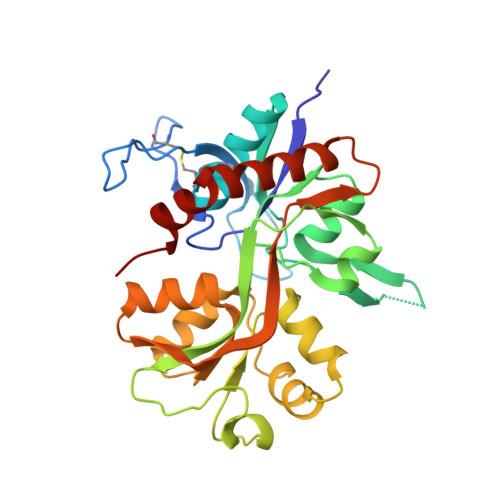
GluN2A-Selective Pyridopyrimidinone Series of NMDAR Positive Allosteric Modulators with an Improved in Vivo Profile.
Villemure, E., Volgraf, M., Jiang, Y., Wu, G., Ly, C.Q., Yuen, P.W., Lu, A., Luo, X., Liu, M., Zhang, S., Lupardus, P.J., Wallweber, H.J., Liederer, B.M., Deshmukh, G., Plise, E., Tay, S., Wang, T.M., Hanson, J.E., Hackos, D.H., Scearce-Levie, K., Schwarz, J.B., Sellers, B.D.(2017) ACS Med Chem Lett 8: 84-89
- PubMed: 28105280 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.6b00388
- Primary Citation Related Structures:
5TP9, 5TPA - PubMed Abstract:
The N -methyl-d-aspartate receptor (NMDAR) is an ionotropic glutamate receptor, gated by the endogenous coagonists glutamate and glycine, permeable to Ca 2+ and Na + . NMDAR dysfunction is associated with numerous neurological and psychiatric disorders, including schizophrenia, depression, and Alzheimer's disease. Recently, we have disclosed GNE-0723 ( 1 ), a GluN2A subunit-selective and brain-penetrant positive allosteric modulator (PAM) of NMDARs. This work highlights the discovery of a related pyridopyrimidinone core with distinct structure-activity relationships, despite the structural similarity to GNE-0723. GNE-5729 ( 13 ), a pyridopyrimidinone-based NMDAR PAM, was identified with both an improved pharmacokinetic profile and increased selectivity against AMPARs. We also include X-ray structure analysis and modeling to propose hypotheses for the activity and selectivity differences.
- Department of Discovery Chemistry, Department of Neurosciences, Department of Biochemical and Cellular Pharmacology, Department of Drug Metabolism and Pharmacokinetics, and Department of Structural Biology, Genentech, Inc. , 1 DNA Way, South San Francisco, California 94080, United States.
Organizational Affiliation: