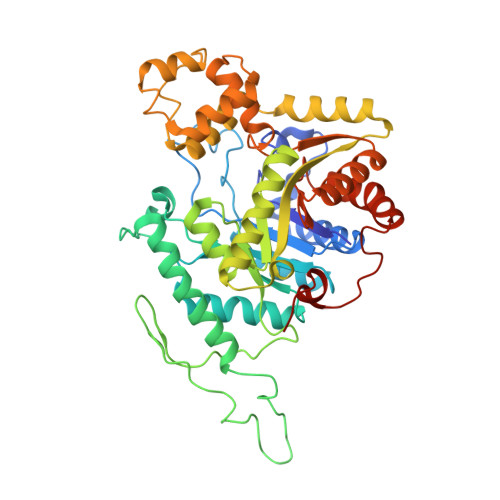
Crystal structure of dibenzothiophene sulfone monooxygenase BdsA from Bacillus subtilis WU-S2B
Okai, M., Lee, W.C., Guan, L.J., Ohshiro, T., Izumi, Y., Tanokura, M.(2017) Proteins 85: 1171-1177
- PubMed: 28205250 Search on PubMed
- DOI: https://doi.org/10.1002/prot.25267
- Primary Citation Related Structures:
5TLC - PubMed Abstract:
The dibenzothiophene (DBT) sulfone monooxygenase BdsA from Bacillus subtilis WU-S2B catalyzes the conversion of DBT sulfone to 2'-hydroxybiphenyl 2-sulfinate. We report the crystal structures of BdsA at a resolution of 2.80 Å. BdsA exists as a homotetramer with a dimer-of-dimers configuration in the crystal, and the interaction between E288 and R296 in BdsA is important for tetramer formation. A structural comparison with homologous proteins shows that the orientation and location of the α9-α12 helices in BdsA are closer to those of the closed form than those of the open form in the EDTA monooxygenase EmoA. Proteins 2017; 85:1171-1177. © 2017 Wiley Periodicals, Inc.
- Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, University of Tokyo, Bunkyo-ku, Tokyo, 113-8657, Japan.
Organizational Affiliation: