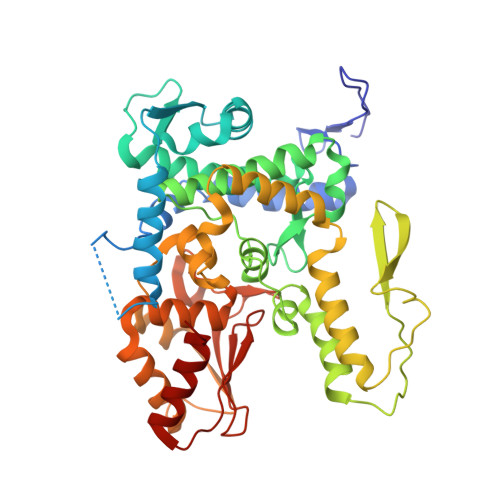
A Tunable Brake for HECT Ubiquitin Ligases.
Chen, Z., Jiang, H., Xu, W., Li, X., Dempsey, D.R., Zhang, X., Devreotes, P., Wolberger, C., Amzel, L.M., Gabelli, S.B., Cole, P.A.(2017) Mol Cell 66: 345-357.e6
- PubMed: 28475870 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2017.03.020
- Primary Citation Related Structures:
5TJ7, 5TJ8, 5TJQ - PubMed Abstract:
The HECT E3 ligases ubiquitinate numerous transcription factors and signaling molecules, and their activity must be tightly controlled to prevent cancer, immune disorders, and other diseases. In this study, we have found unexpectedly that peptide linkers tethering WW domains in several HECT family members are key regulatory elements of their catalytic activities. Biochemical, structural, and cellular analyses have revealed that the linkers can lock the HECT domain in an inactive conformation and block the proposed allosteric ubiquitin binding site. Such linker-mediated autoinhibition of the HECT domain can be relieved by linker post-translational modifications, but complete removal of the brake can induce hyperactive autoubiquitination and E3 self destruction. These results clarify the mechanisms of several HECT protein cancer associated mutations and provide a new framework for understanding how HECT ubiquitin ligases must be finely tuned to ensure normal cellular behavior.
- Department of Pharmacology and Molecular Sciences, John Hopkins School of Medicine, Baltimore, MD 21205, USA.
Organizational Affiliation: