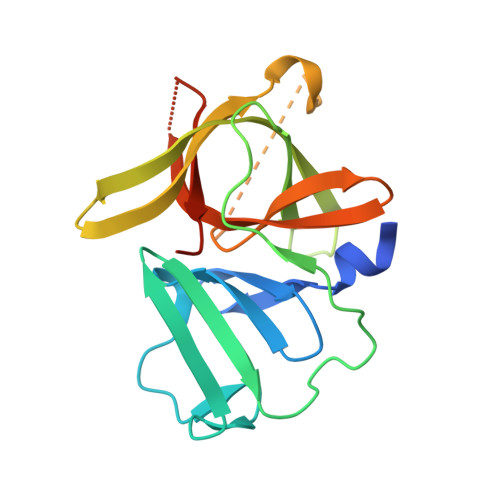
Design, synthesis, and evaluation of a novel series of macrocyclic inhibitors of norovirus 3CL protease.
Damalanka, V.C., Kim, Y., Galasiti Kankanamalage, A.C., Lushington, G.H., Mehzabeen, N., Battaile, K.P., Lovell, S., Chang, K.O., Groutas, W.C.(2016) Eur J Med Chem 127: 41-61
- PubMed: 28038326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ejmech.2016.12.033
- Primary Citation Related Structures:
5TG1, 5TG2 - PubMed Abstract:
Norovirus infections have a major impact on public health worldwide, yet there is a current dearth of norovirus-specific therapeutics and prophylactics. This report describes the discovery of a novel class of macrocyclic inhibitors of norovirus 3C-like protease, a cysteine protease that is essential for virus replication. SAR, structural, and biochemical studies were carried out to ascertain the effect of structure on pharmacological activity and permeability. Insights gained from these studies have laid a solid foundation for capitalizing on the therapeutic potential of the series of inhibitors described herein.
- Department of Chemistry, Wichita State University, Wichita, KS 67260, USA.
Organizational Affiliation: