Lipid interactions and angle of approach to the HIV-1 viral membrane of broadly neutralizing antibody 10E8: Insights for vaccine and therapeutic design.
Irimia, A., Serra, A.M., Sarkar, A., Jacak, R., Kalyuzhniy, O., Sok, D., Saye-Francisco, K.L., Schiffner, T., Tingle, R., Kubitz, M., Adachi, Y., Stanfield, R.L., Deller, M.C., Burton, D.R., Schief, W.R., Wilson, I.A.(2017) PLoS Pathog 13: e1006212-e1006212
- PubMed: 28225819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1006212
- Primary Citation Related Structures:
5SY8, 5T29, 5T5B, 5T6L, 5T80, 5T85, 5TFW - PubMed Abstract:
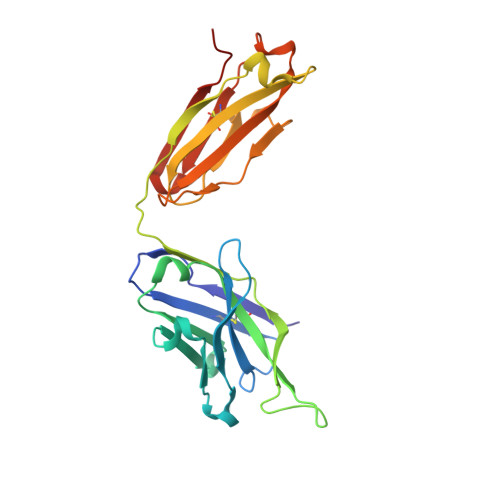
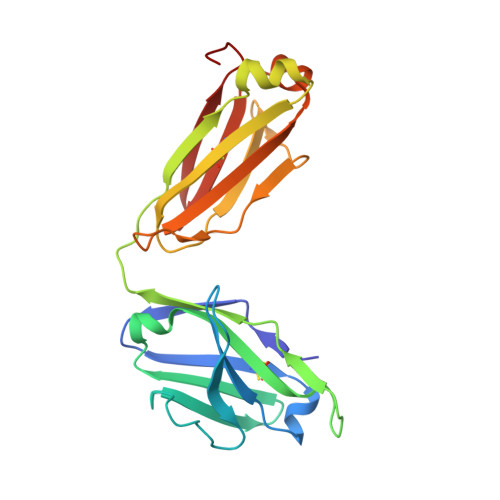
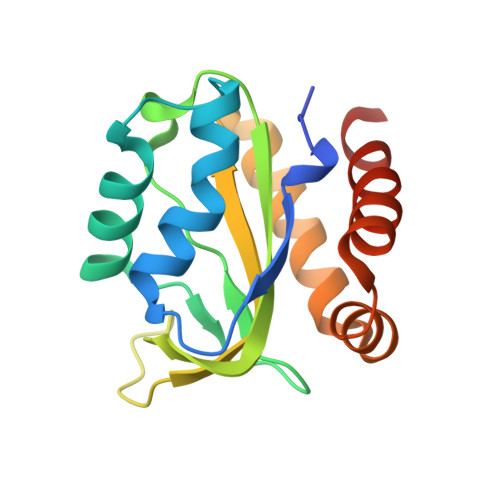
Among broadly neutralizing antibodies to HIV, 10E8 exhibits greater neutralizing breadth than most. Consequently, this antibody is the focus of prophylactic/therapeutic development. The 10E8 epitope has been identified as the conserved membrane proximal external region (MPER) of gp41 subunit of the envelope (Env) viral glycoprotein and is a major vaccine target. However, the MPER is proximal to the viral membrane and may be laterally inserted into the membrane in the Env prefusion form. Nevertheless, 10E8 has not been reported to have significant lipid-binding reactivity. Here we report x-ray structures of lipid complexes with 10E8 and a scaffolded MPER construct and mutagenesis studies that provide evidence that the 10E8 epitope is composed of both MPER and lipid. 10E8 engages lipids through a specific lipid head group interaction site and a basic and polar surface on the light chain. In the model that we constructed, the MPER would then be essentially perpendicular to the virion membrane during 10E8 neutralization of HIV-1. As the viral membrane likely also plays a role in selecting for the germline antibody as well as size and residue composition of MPER antibody complementarity determining regions, the identification of lipid interaction sites and the MPER orientation with regard to the viral membrane surface during 10E8 engagement can be of great utility for immunogen and therapeutic design.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, California, United States of America.
Organizational Affiliation: