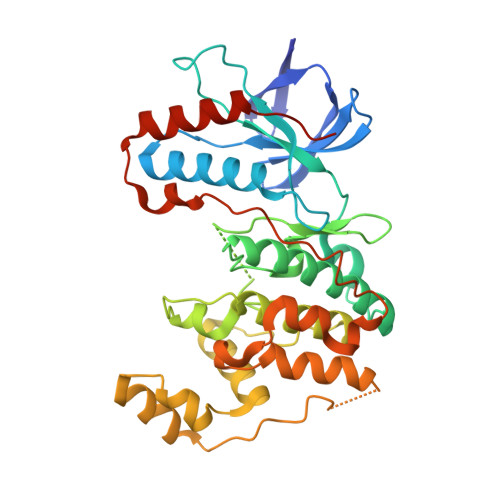
Optimized Target Residence Time: Type I1/2 Inhibitors for p38 alpha MAP Kinase with Improved Binding Kinetics through Direct Interaction with the R-Spine.
Wentsch, H.K., Walter, N.M., Buhrmann, M., Mayer-Wrangowski, S., Rauh, D., Zaman, G.J.R., Willemsen-Seegers, N., Buijsman, R.C., Henning, M., Dauch, D., Zender, L., Laufer, S.(2017) Angew Chem Int Ed Engl 56: 5363-5367
- PubMed: 28397331 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201701185
- Primary Citation Related Structures:
5TBE, 5TCO - PubMed Abstract:
Skepinone-L was recently reported to be a p38α MAP kinase inhibitor with high potency and excellent selectivity in vitro and in vivo. However, this class of compounds still act as fully ATP-competitive Type I binders which, furthermore, suffer from short residence times at the enzyme. We herein describe a further development with the first Type I1/2 binders for p38α MAP kinase. Type I1/2 inhibitors interfere with the R-spine, inducing a glycine flip and occupying both hydrophobic regions I and II. This design approach leads to prolonged target residence time, binding to both the active and inactive states of the kinase, excellent selectivity, excellent potency on the enzyme level, and low nanomolar activity in a human whole blood assay. This promising binding mode is proven by X-ray crystallography.
- Institute of Pharmaceutical Sciences, Pharmaceutical and Medicinal Chemistry, Eberhard Karls Universität Tübingen, Auf der Morgenstelle 8, 72076, Tübingen, Germany.
Organizational Affiliation: