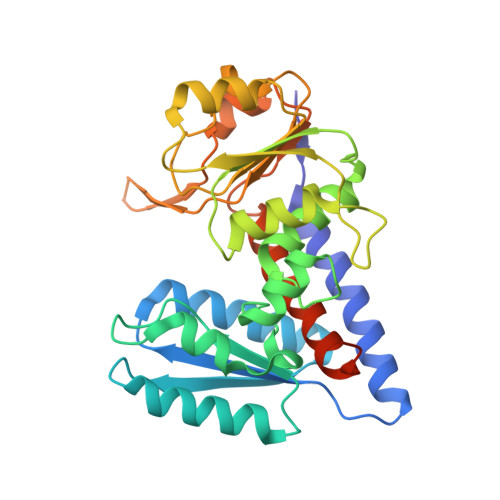
Crystal Structure of the Emerging Cancer Target MTHFD2 in Complex with a Substrate-Based Inhibitor.
Gustafsson, R., Jemth, A.S., Gustafsson, N.M., Farnegardh, K., Loseva, O., Wiita, E., Bonagas, N., Dahllund, L., Llona-Minguez, S., Haggblad, M., Henriksson, M., Andersson, Y., Homan, E., Helleday, T., Stenmark, P.(2017) Cancer Res 77: 937-948
- PubMed: 27899380 Search on PubMed
- DOI: https://doi.org/10.1158/0008-5472.CAN-16-1476
- Primary Citation Related Structures:
5TC4 - PubMed Abstract:
To sustain their proliferation, cancer cells become dependent on one-carbon metabolism to support purine and thymidylate synthesis. Indeed, one of the most highly upregulated enzymes during neoplastic transformation is MTHFD2, a mitochondrial methylenetetrahydrofolate dehydrogenase and cyclohydrolase involved in one-carbon metabolism. Because MTHFD2 is expressed normally only during embryonic development, it offers a disease-selective therapeutic target for eradicating cancer cells while sparing healthy cells. Here we report the synthesis and preclinical characterization of the first inhibitor of human MTHFD2. We also disclose the first crystal structure of MTHFD2 in complex with a substrate-based inhibitor and the enzyme cofactors NAD + and inorganic phosphate. Our work provides a rationale for continued development of a structural framework for the generation of potent and selective MTHFD2 inhibitors for cancer treatment. Cancer Res; 77(4); 937-48. ©2017 AACR .
- Department of Biochemistry and Biophysics, Stockholm University, Stockholm, Sweden.
Organizational Affiliation: