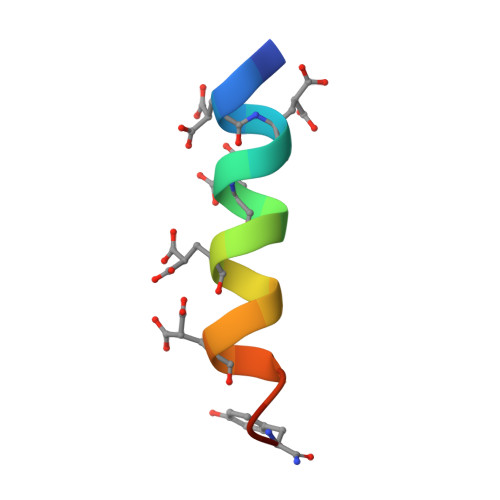
Discerning the Role of the Hydroxyproline Residue in the Structure of Conantokin Rl-B and Its Role in GluN2B Subunit-Selective Antagonistic Activity toward N-Methyl-d-Aspartate Receptors.
Yuan, Y., Balsara, R.D., Zajicek, J., Kunda, S., Castellino, F.J.(2016) Biochemistry 55: 7112-7122
- PubMed: 27981829 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.6b00962
- Primary Citation Related Structures:
5TBG, 5TBQ, 5TBR - PubMed Abstract:
Conantokins (con) are short γ-carboxyglutamate (Gla)-containing polypeptides expressed by marine snails that function as antagonists of N-methyl-d-aspartate receptor (NMDAR) ion channels. The Gla residues govern structural conformations and antagonistic activities of the conantokins. In addition to Gla, some conantokins, e.g., conRl-B, also contain a hydroxyproline (HyP or O) residue, which in this case is centrally located in the peptide at position 10. Because conRl-B specifically inhibits ion channels of GluN2B subunit-containing heterotetrameric NMDARs, we evaluated the unusual role of HyP 10 in this effect. To accomplish this goal, we examined synthetic variants of conRl-B in which HyP 10 was either deleted (conRl-B[ΔO 10 ]) or replaced with alanine (conRl-B[O 10 A]) or proline (conRl-B[O 10 P]). The solution structures of these variants were determined by nuclear magnetic resonance spectroscopy. Deletion of HyP 10 , or replacement of HyP 10 with Ala 10 , attenuated the distortion in the central region of the apo-conRl-B helix and allowed Mg 2+ -complexed end-to-end α-helix formation. The inhibitory properties of these variants were assessed by measuring NMDA/Gly-stimulated intracellular Ca 2+ influx in mice neurons. ConRl-B[O 10 P] retained its NMDAR ion channel inhibitory activity in wild-type (WT) neurons but lost its GluN2B specificity, whereas conRl-B[ΔO 10 ] showed overall diminished inhibitory function. ConRl-B[O 10 A] showed attenuated inhibitory function but retained its GluN2B specificity. Thus, HyP 10 plays a critical role in maintaining the structural integrity of conRl-B, which can be correlated with its GluN2B subunit-selective inhibition. Weakened inhibition by conRl-B was also observed in neurons lacking either the GluN2C or GluN2D subunit, compared to WT neurons. This suggests that GluN2C and GluN2D are also required for inhibition by conRl-B.
- W. M. Keck Center for Transgene Research and ‡Department of Chemistry and Biochemistry, University of Notre Dame , Notre Dame, Indiana 46556, United States.
Organizational Affiliation: