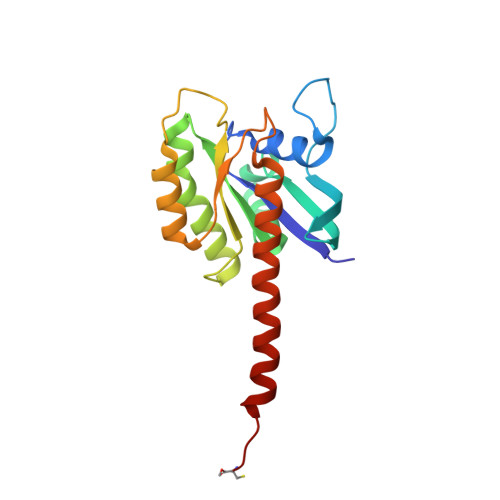
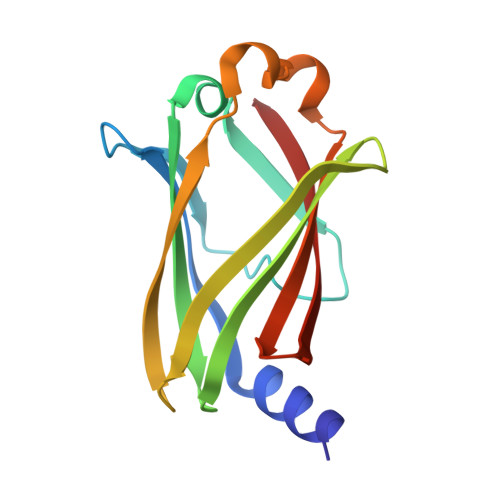
Structural basis of recognition of farnesylated and methylated KRAS4b by PDE delta.
Dharmaiah, S., Bindu, L., Tran, T.H., Gillette, W.K., Frank, P.H., Ghirlando, R., Nissley, D.V., Esposito, D., McCormick, F., Stephen, A.G., Simanshu, D.K.(2016) Proc Natl Acad Sci U S A 113: E6766-E6775
- PubMed: 27791178 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1615316113
- Primary Citation Related Structures:
5TAR, 5TB5 - PubMed Abstract:
Farnesylation and carboxymethylation of KRAS4b (Kirsten rat sarcoma isoform 4b) are essential for its interaction with the plasma membrane where KRAS-mediated signaling events occur. Phosphodiesterase-δ (PDEδ) binds to KRAS4b and plays an important role in targeting it to cellular membranes. We solved structures of human farnesylated-methylated KRAS4b in complex with PDEδ in two different crystal forms. In these structures, the interaction is driven by the C-terminal amino acids together with the farnesylated and methylated C185 of KRAS4b that binds tightly in the central hydrophobic pocket present in PDEδ. In crystal form II, we see the full-length structure of farnesylated-methylated KRAS4b, including the hypervariable region. Crystal form I reveals structural details of farnesylated-methylated KRAS4b binding to PDEδ, and crystal form II suggests the potential binding mode of geranylgeranylated-methylated KRAS4b to PDEδ. We identified a 5-aa-long sequence motif (Lys-Ser-Lys-Thr-Lys) in KRAS4b that may enable PDEδ to bind both forms of prenylated KRAS4b. Structure and sequence analysis of various prenylated proteins that have been previously tested for binding to PDEδ provides a rationale for why some prenylated proteins, such as KRAS4a, RalA, RalB, and Rac1, do not bind to PDEδ. Comparison of all four available structures of PDEδ complexed with various prenylated proteins/peptides shows the presence of additional interactions due to a larger protein-protein interaction interface in KRAS4b-PDEδ complex. This interface might be exploited for designing an inhibitor with minimal off-target effects.
- NCI RAS Initiative, Cancer Research Technology Program, Frederick National Laboratory for Cancer Research, Leidos Biomedical Research, Inc., Frederick, MD 21701.
Organizational Affiliation: