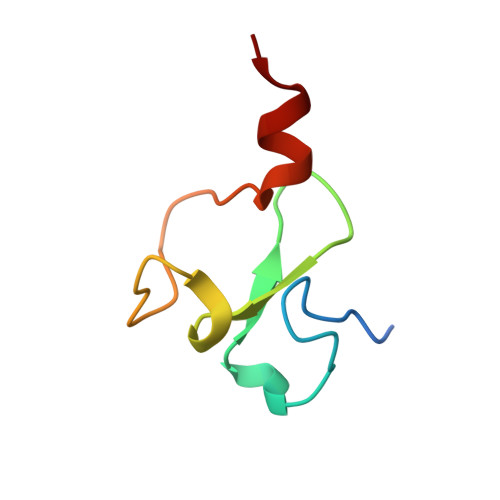
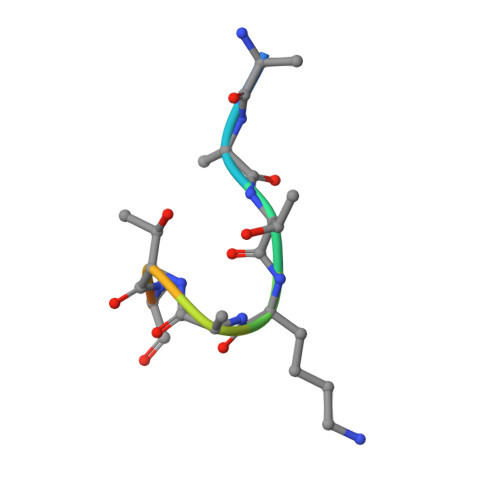
Structural basis of molecular recognition of helical histone H3 tail by PHD finger domains.
Bortoluzzi, A., Amato, A., Lucas, X., Blank, M., Ciulli, A.(2017) Biochem J 474: 1633-1651
- PubMed: 28341809 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20161053
- Primary Citation Related Structures:
5T8R - PubMed Abstract:
The plant homeodomain (PHD) fingers are among the largest family of epigenetic domains, first characterized as readers of methylated H3K4. Readout of histone post-translational modifications by PHDs has been the subject of intense investigation; however, less is known about the recognition of secondary structure features within the histone tail itself. We solved the crystal structure of the PHD finger of the bromodomain adjacent to zinc finger 2A [BAZ2A, also known as TIP5 (TTF-I/interacting protein 5)] in complex with unmodified N-terminal histone H3 tail. The peptide is bound in a helical folded-back conformation after K4, induced by an acidic patch on the protein surface that prevents peptide binding in an extended conformation. Structural bioinformatics analyses identify a conserved Asp/Glu residue that we name 'acidic wall', found to be mutually exclusive with the conserved Trp for K4Me recognition. Neutralization or inversion of the charges at the acidic wall patch in BAZ2A, and homologous BAZ2B, weakened H3 binding. We identify simple mutations on H3 that strikingly enhance or reduce binding, as a result of their stabilization or destabilization of H3 helicity. Our work unravels the structural basis for binding of the helical H3 tail by PHD fingers and suggests that molecular recognition of secondary structure motifs within histone tails could represent an additional layer of regulation in epigenetic processes.
- Division of Biological Chemistry and Drug Discovery, School of Life Sciences, University of Dundee, James Black Centre, Dow Street, Dundee DD1 5EH, U.K.
Organizational Affiliation: