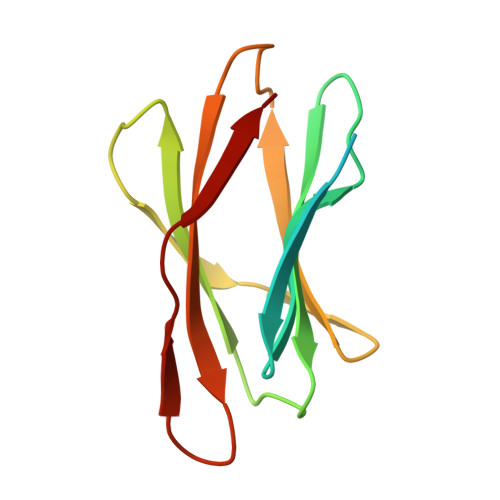
Properties of a family 56 carbohydrate-binding module and its role in the recognition and hydrolysis of beta-1,3-glucan.
Hettle, A., Fillo, A., Abe, K., Massel, P., Pluvinage, B., Langelaan, D.N., Smith, S.P., Boraston, A.B.(2017) J Biological Chem 292: 16955-16968
- PubMed: 28827308 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.806711
- Primary Citation Related Structures:
5T7A - PubMed Abstract:
BH0236 from Bacillus halodurans is a multimodular β-1,3-glucanase comprising an N-terminal family 81 glycoside hydrolase catalytic module, an internal family 6 carbohydrate-binding module (CBM) that binds the nonreducing end of β-1,3-glucan chains, and an uncharacterized C-terminal module classified into CBM family 56. Here, we determined that this latter CBM, BhCBM56, bound the soluble β-1,3-glucan laminarin with a dissociation constant ( K d ) of ∼26 μm and displayed higher affinity for insoluble β-1,3-glucans with K d values of ∼2-10 μm but lacked affinity for β-1,3-glucooligosaccharides. The X-ray crystal structure of BhCBM56 and NMR-derived chemical shift mapping of the binding site revealed a β-sandwich fold, with the face of one β-sheet possessing the β-1,3-glucan-binding surface. On the basis of the functional and structural properties of BhCBM56, we propose that it binds a quaternary polysaccharide structure, most likely the triple helix adopted by polymerized β-1,3-glucans. Consistent with the BhCBM56 and BhCBM6/56 binding profiles, deletion of the CBM56 from BH0236 decreased activity of the enzyme on the insoluble β-1,3-glucan curdlan but not on soluble laminarin; additional deletion of the CBM6 also did not affect laminarin degradation but further decreased curdlan hydrolysis. The pseudo-atomic solution structure of BH0236 determined by small-angle X-ray scattering revealed structural insights into the nature of avid binding by the BhCBM6/56 pair and how the orientation of the active site in the catalytic module factors into recognition and degradation of β-1,3-glucans. Our findings reinforce the notion that catalytic modules and their cognate CBMs have complementary specificities, including targeting of polysaccharide quaternary structure.
- From the Department of Biochemistry and Microbiology, University of Victoria, Victoria, British Columbia V8W 3P6, Canada and.
Organizational Affiliation: